Session Information
Date: Friday, November 6, 2020
Title: Miscellaneous Rheumatic & Inflammatory Diseases Poster I: Diagnosis and Testing
Session Type: Poster Session A
Session Time: 9:00AM-11:00AM
Background/Purpose: Monogenic autoinflammatory diseases (AID) are caused by mutations in innate immune signaling genes. The effect of these mutations on the risk of allergy is unknown. We previously reported results of our preliminary analysis, which indicated that allergy-associated features were rare in AID based on retrospective review [1]. We have now performed a full systematic analysis using a combination of retrospective chart review, standardized questionnaires, and T cell immunophenotyping. This revealed common and disease-specific correlations with allergy-associated clinical and immunologic phenotypes. This study was the first of its kind to investigate the prevalence of allergic features in multiple monogenic AID, and the first to do so within the U.S. population.
Methods: We investigated the prevalence of allergic clinical and immunologic phenotypes in AID caused by pathogenic mutations in MEFV, TNFRSF1A, MVK, NLRP3, PSTPIP1, proteasome genes, STING, TNFAIP3, and ADA2. We used retrospective chart review and questionnaires to assess clinical and laboratory features of patients enrolled on NIH protocol 94-HG-0105 or 17-I-0016. Questionnaires were developed by a panel of clinical allergists, clinical rheumatologists, and survey researchers. Six clinical syndromes were assessed (Table 1). Surveys were administered via the telephone, internet, and in the NIH Clinical Center. Informed consent was obtained from all participants. Comparator data was obtained from the National Center for Health Statistics and Chicago Food Allergy Research Surveys [2-6]. To determine if T helper 2 (Th2)/Th9 cells were expanded in AID, we stimulated peripheral blood mononuclear cells (PBMCs) and analyzed the data with flow cytometry.
Results: Allergic rhinitis, eczema, and eosinophilic gastrointestinal (GI) inflammation were prevalent in multiple AID (Table 1). Asthma was rare in patients with MEFV and proteasome mutations but common in patients with NLRP3 mutations. A trend towards increased food allergy prevalence was seen in patients with NLRP3, TNFAIP3, and proteasome mutations. Median IgE was reduced in patients with ADA2 and MVK mutations; median absolute eosinophil count (AEC) was reduced in patients with most AID but elevated in patients with NLRP3 mutations (Figure 1). T helper 2 (Th2) cells were expanded in patients with NLRP3, TNFRSF1A, and MVK mutations; Th9 cells were expanded in patients with TNFAIP3 mutations (Figure 2).
Conclusion: Allergy-associated clinical and immunological phenotypes are prevalent in many AID, reinforcing a role for innate immunity in allergy pathogenesis. Further investigations are needed to determine the role of allergy-associated factors in AID pathology.
References
- Dizon B, et al. Prevalence of Atopic Features in Classic Autoinflammatory Diseases. 2019 ACR Annual Meeting.
- National Center for Health Statistics.
- Gupta RS, et al. The Public Health Impact of Parent-Reported Childhood Food Allergies in the United States. Pediatrics 2018; 142.
- Gupta RS, Warren CM, Smith BM, Jiang J, Blumenstock JA, Davis MM, et al. Prevalence and Severity of Food Allergies Among US Adults. JAMA Netw Open 2019; 2:e185630.
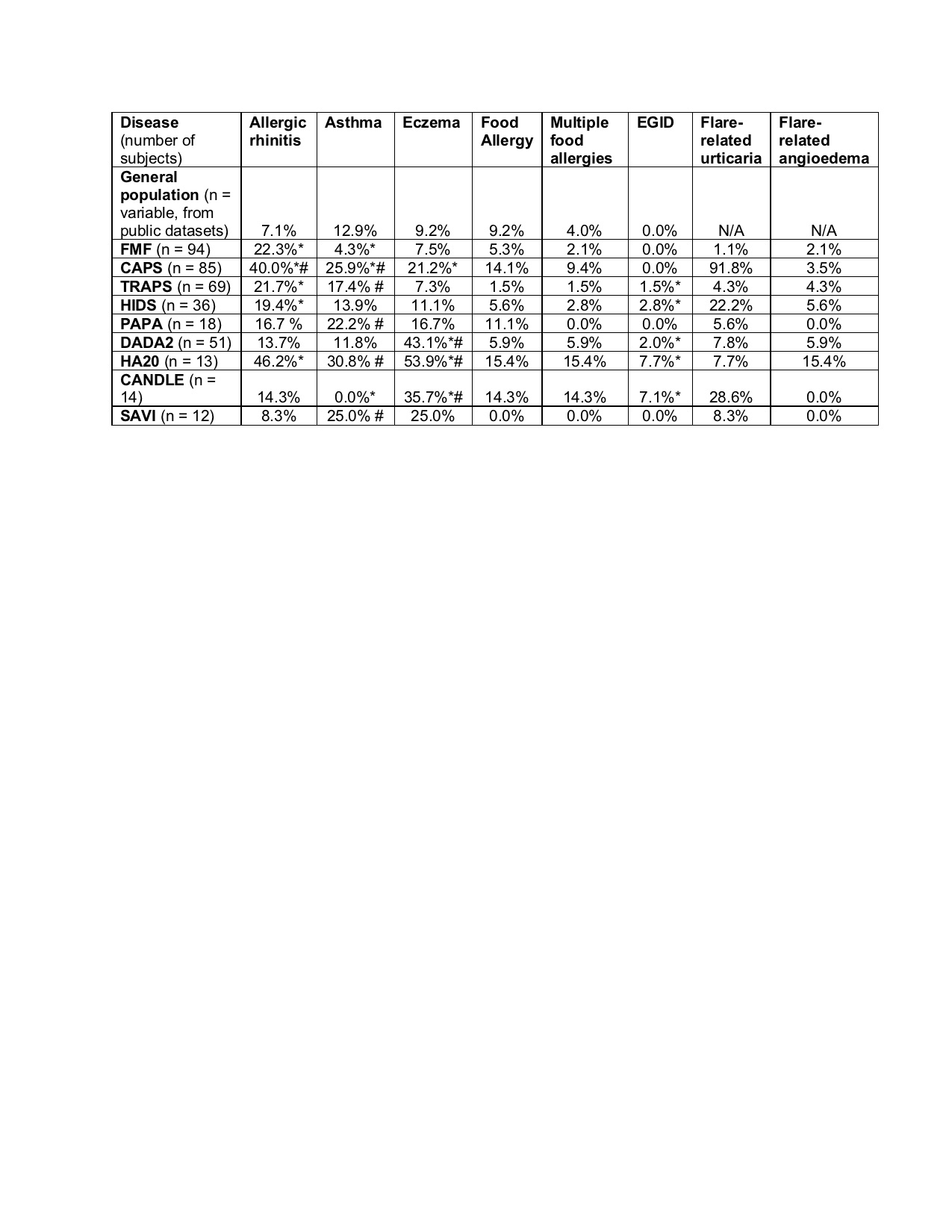 Table 1. Prevalence of atopic features in patients with autoinflammatory disease confirmed by genetic testing, and comparison data in the general population. Column shows prevalence in each population. FMF = Familial Mediterranean Fever; CAPS = Cryopyrin-Associated Periodic Syndrome; TRAPS = Tumor necrosis factor Receptor-Associated Periodic Syndrome (TRAPS); DADA2 = Deficiency of Adenosine Deaminase 2; HIDS = Hyper-IgD Syndrome; PAPA = Pyogenic Arthritis, Pyoderma gangrenosum, and Acne; HA20 = Haploinsufficiency of A20; CANDLE = Chronic Atypical Neutrophilic Dermatosis with Lipodystrophy and Elevated temperature; SAVI = STING-Associated Vasculopathy with onset in Infancy. *FDR < 0.05 vs. general US population. #FDR < 0.05 vs. FMF; Benjamini-Hochberg adjusted Fisher t-test.
Table 1. Prevalence of atopic features in patients with autoinflammatory disease confirmed by genetic testing, and comparison data in the general population. Column shows prevalence in each population. FMF = Familial Mediterranean Fever; CAPS = Cryopyrin-Associated Periodic Syndrome; TRAPS = Tumor necrosis factor Receptor-Associated Periodic Syndrome (TRAPS); DADA2 = Deficiency of Adenosine Deaminase 2; HIDS = Hyper-IgD Syndrome; PAPA = Pyogenic Arthritis, Pyoderma gangrenosum, and Acne; HA20 = Haploinsufficiency of A20; CANDLE = Chronic Atypical Neutrophilic Dermatosis with Lipodystrophy and Elevated temperature; SAVI = STING-Associated Vasculopathy with onset in Infancy. *FDR < 0.05 vs. general US population. #FDR < 0.05 vs. FMF; Benjamini-Hochberg adjusted Fisher t-test.
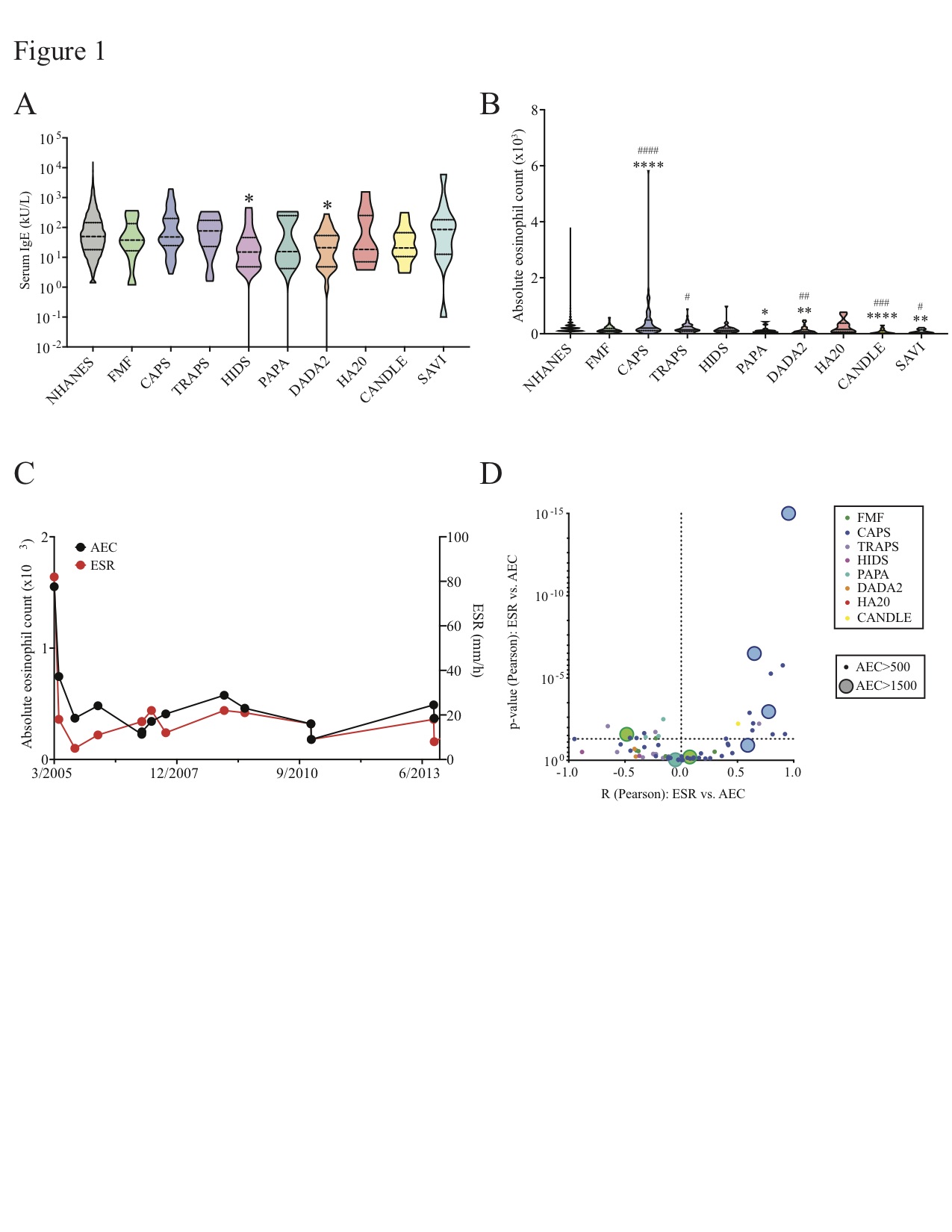 Figure 1: Eosinophilia and absolute eosinophil counts (AEC) in the general US population and in nine monogenic autoinflammatory diseases (AID). Data are shown for the general US population, Familial Mediterranean Fever (FMF); Cryopyrin-Associated Periodic Syndrome (CAPS); TNF-Receptor Associated Periodic Syndrome (TRAPS); Hyper-IgD Syndrome (HIDS); Pyogenic Arthritis, Pyoderma gangrenosum, and Acne (PAPA); Deficiency of ADA2 (DADA2); Haploinsufficiency of A20 (HA20); Chronic Atypical Neutrophilic Dermatosis with Lipodystrophy and Elevated temperature (CANDLE); and STING Associated Vasculopathy with onset in Infancy (SAVI). (A) Violin plot shows serum IgE levels for each population. (B) Violin plot shows serum IgE levels for each population. (C) Representative plots are shown for one CAPS patient with hypereosinophilia: dot plot of AEC and ESR values over a period of 8 years. (D) Volcano plot shows correlation (R, Pearson test) vs. p-value (Pearson test) of AEC vs. erythrocyte sedimentation rate (ESR, mm/h) for each AID patient with eosinophilia (AEC > 500). Colors of dots indicate specific AID (FMF, green; CAPS, dark blue; TRAPS, purple; HIDS, fuchsia; PAPA, teal; DADA2, orange; HA20, red; CANDLE, yellow). Size of dots distinguishes patients with eosinophilia (AEC > 500) from those with hypereosinophilia (AEC > 1500). *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.001 vs. general US population. #p < 0.05, ##p < 0.01, ###p < 0.005, ###p < 0.001 vs. FMF; Benjamini-Hochberg adjusted Mann-Whitney (B).
Figure 1: Eosinophilia and absolute eosinophil counts (AEC) in the general US population and in nine monogenic autoinflammatory diseases (AID). Data are shown for the general US population, Familial Mediterranean Fever (FMF); Cryopyrin-Associated Periodic Syndrome (CAPS); TNF-Receptor Associated Periodic Syndrome (TRAPS); Hyper-IgD Syndrome (HIDS); Pyogenic Arthritis, Pyoderma gangrenosum, and Acne (PAPA); Deficiency of ADA2 (DADA2); Haploinsufficiency of A20 (HA20); Chronic Atypical Neutrophilic Dermatosis with Lipodystrophy and Elevated temperature (CANDLE); and STING Associated Vasculopathy with onset in Infancy (SAVI). (A) Violin plot shows serum IgE levels for each population. (B) Violin plot shows serum IgE levels for each population. (C) Representative plots are shown for one CAPS patient with hypereosinophilia: dot plot of AEC and ESR values over a period of 8 years. (D) Volcano plot shows correlation (R, Pearson test) vs. p-value (Pearson test) of AEC vs. erythrocyte sedimentation rate (ESR, mm/h) for each AID patient with eosinophilia (AEC > 500). Colors of dots indicate specific AID (FMF, green; CAPS, dark blue; TRAPS, purple; HIDS, fuchsia; PAPA, teal; DADA2, orange; HA20, red; CANDLE, yellow). Size of dots distinguishes patients with eosinophilia (AEC > 500) from those with hypereosinophilia (AEC > 1500). *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.001 vs. general US population. #p < 0.05, ##p < 0.01, ###p < 0.005, ###p < 0.001 vs. FMF; Benjamini-Hochberg adjusted Mann-Whitney (B).
 Figure 2. T helper immunophenotypes in seven monogenic autoinflammatory diseases (AID). Data are shown for healthy volunteers (HV); atopic dermatitis patients; Familial Mediterranean Fever (FMF); Cryopyrin-Associated Periodic Syndrome (CAPS); TNF-Receptor Associated Periodic Syndrome (TRAPS); Hyper-IgD Syndrome (HIDS); Pyogenic Arthritis, Pyoderma gangrenosum, and Acne (PAPA); Deficiency of ADA2 (DADA2); and Haploinsufficiency of A20 (HA20). Violin plots show percentage of memory T helper cells (CD3+ CD4+ CD8- CD45RO+) producing IL-17A (A), IFN-gamma (B), IL-9 (C), IL-13 (D), IL-4 (E), and IL-5 (F). *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.001 vs. HC. # < 0.05, ##p < 0.01, ###p < 0.005, ###p < 0.001 vs. FMF; Benjamini-Hochberg adjusted Mann Whitney.
Figure 2. T helper immunophenotypes in seven monogenic autoinflammatory diseases (AID). Data are shown for healthy volunteers (HV); atopic dermatitis patients; Familial Mediterranean Fever (FMF); Cryopyrin-Associated Periodic Syndrome (CAPS); TNF-Receptor Associated Periodic Syndrome (TRAPS); Hyper-IgD Syndrome (HIDS); Pyogenic Arthritis, Pyoderma gangrenosum, and Acne (PAPA); Deficiency of ADA2 (DADA2); and Haploinsufficiency of A20 (HA20). Violin plots show percentage of memory T helper cells (CD3+ CD4+ CD8- CD45RO+) producing IL-17A (A), IFN-gamma (B), IL-9 (C), IL-13 (D), IL-4 (E), and IL-5 (F). *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.001 vs. HC. # < 0.05, ##p < 0.01, ###p < 0.005, ###p < 0.001 vs. FMF; Benjamini-Hochberg adjusted Mann Whitney.
To cite this abstract in AMA style:
Kitakule M, Dizon B, Gutierrez-Huerta C, Blackstone S, Burma A, Son A, Deuitch N, Rosenzweig S, Komarow H, Stone D, Jones A, Nehrbecky M, Hoffmann P, Romeo T, de Jesus A, Alehashemi S, Garg M, Torreggiani S, Montealegre Sanchez G, Honer K, Barron K, Aksentijevich I, Ombrello A, Goldbach-Mansky R, Kastner D, Milner J, Frischmeyer-Guerrerio P, Schwartz D. Systematic Evaluation of Nine Monogenic Autoinflammatory Diseases Reveals Common and Disease-Specific Correlations with Allergy-Associated Features [abstract]. Arthritis Rheumatol. 2020; 72 (suppl 10). https://acrabstracts.org/abstract/systematic-evaluation-of-nine-monogenic-autoinflammatory-diseases-reveals-common-and-disease-specific-correlations-with-allergy-associated-features/. Accessed .« Back to ACR Convergence 2020
ACR Meeting Abstracts - https://acrabstracts.org/abstract/systematic-evaluation-of-nine-monogenic-autoinflammatory-diseases-reveals-common-and-disease-specific-correlations-with-allergy-associated-features/
