Session Information
Session Type: Abstract Session
Session Time: 10:00AM-10:50AM
Background/Purpose: A new dimension has been added to the link between diet and health, the gut microbiome. Of particular interest is the influence of diet on the microbiome and how this affects pain/inflammation by modifying circulating pro/anti-inflammatory metabolites. Microbiome-derived metabolites have been related to beneficial or deleterious effects on the host. Thus, changes in diet and microbiome might change the systemic metabolite profiling and influence pain and inflammation in rheumatoid arthritis (RA). We recently described that RA patients exposed to an anti-inflammatory ITIS diet had improvement of clinical outcomes. Here, we examine the effect of the ITIS diet on gut microbiome and fecal and plasma metabolome in RA patients.
Methods: We conducted an open-label pilot trial to evaluate feasibility and efficacy of a 2-week anti-inflammatory diet (ITIS) in RA patients. 17 patients with active RA (at least 3 tender and 3 swollen joints) participated. Physical examination and collection of fecal and plasma samples for microbiome and metabolomics were performed at each visit. Using a 50% pain improvement, patients were categorized as responders (R, N=11) or non-responders (NR, N=6). 16S rRNA gene amplicon profiling of the stools and untargeted mass spectrometry (MS)-based metabolomics in stool and plasma were performed. Dietary adherence was evaluated based on food logs and metabolomics readouts.
Results: Clinical outcomes showed significant improvements (pain: 3.9±2.3 before vs 2.45±2.4 after diet, p< 0.01; Clinical Disease Activity Index (CDAI): (29±11.7 before vs 12.7±11.3 after diet, p< 0.001). A diet score (up to 200) showed good adherence, with an average improvement of 148.18 (± 58.10). Changes due to diet intervention were captured by MS analysis and reflected the prescribed diet as well as the individual dietary diary entries (Figure 1A). Alpha diversity was higher at baseline in R (Figure 1B). Interestingly, R had a smaller change in alpha diversity after diet than NR suggesting that an already high alpha diversity with relatively limited capacity for further change may be necessary for response with this dietary intervention (Figure 1C). Ruminococcaceae, Clostridium, Bacteroides and Lachnospira genera negatively correlated with measures of disease activity at baseline, while Enterobacteriaceae positively correlated with the same measures. (Figure 1D). Baseline plasma metabolites profiles were different in R compared to NR (Figure 2A). The differential analysis showed that responders of both Pain 50 and CDAI 70 were enriched for both conjugated linoleic acid (CLA) and Mollicutes or depleted in taurocholic acid, phosphocholines and Coriobacteriales (Figure 2B-C). Of interes, CLA is an anti-inflammatory metabolite previously associated with dietary changes in the number of types of plants eaten.
Conclusion: The “ITIS” diet induced changes in both metabolome and microbiome which correlate with clinical outcome measures. Further studies are needed to provide insight into specific long-term dietary interventions or supplements as a complementary treatment in RA, and to establish the scientific basis for using diet to adjust the gut microbiome to improve RA management.
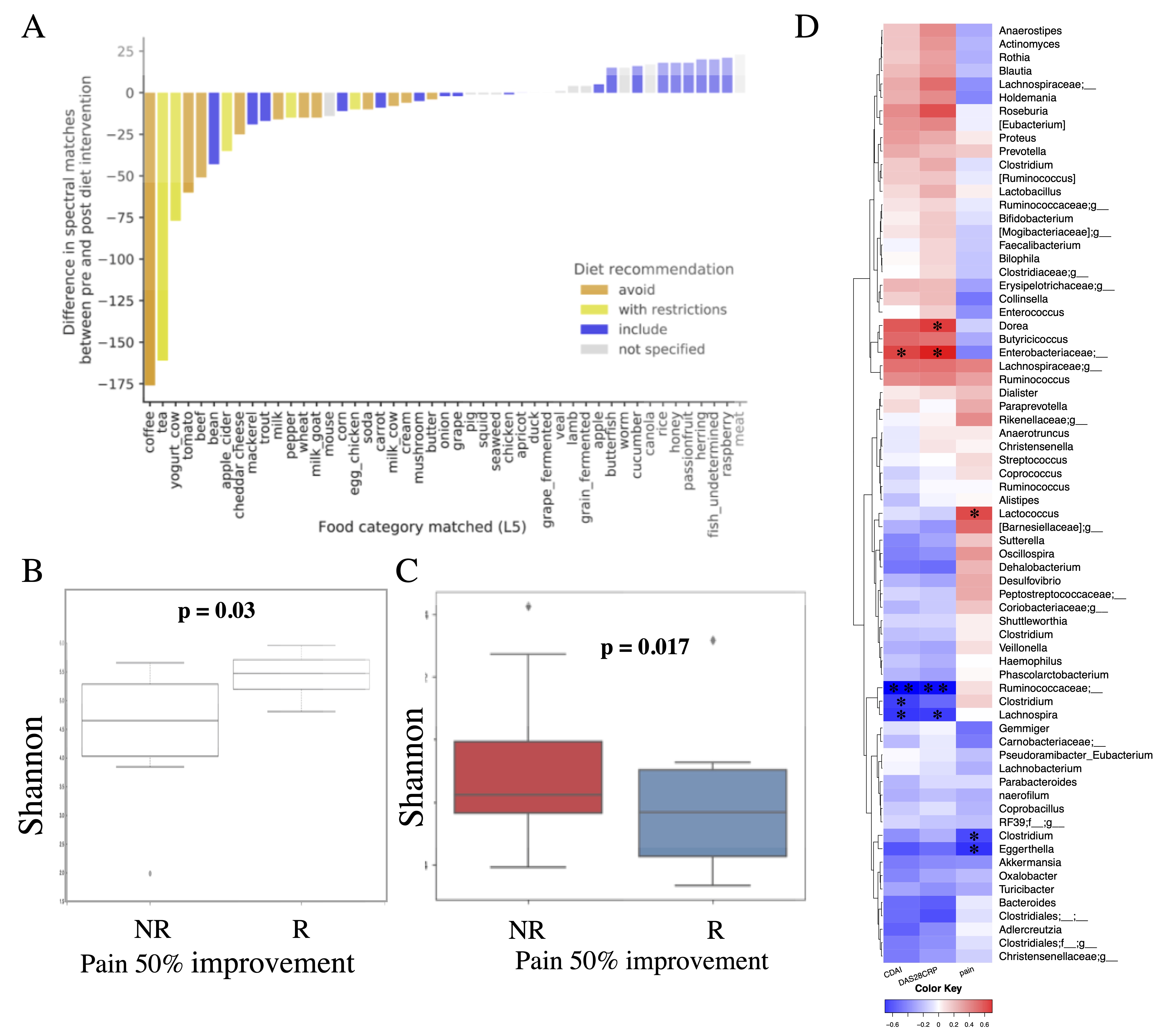 Figure 1. Adherence to diet and microbiome at baseline. A) Adherence to diet evaluated by the plasma metabolomics readout. B) Alpha diversity (Shannon) at baseline is significantly higher in patients with at least 50% pain improvement (R) compared to patients with a less than 50% improvement in pain (NR). C) Change in alpha diversity from day-14 (before diet) to day +14 (after diet) in R versus NR, assessed by at least 50% pain improvement. Boxes represent group means with standard deviations. D) Correlation (Spearman rank correlation) of bacterial genus with clinical outcomes at baseline (CDAI – clinical disease activity index, DAS28CRP – disease activity score using 28 joint count and C reactive protein). The significant correlations are marked with * (p < 0.05) or ** (p < 0.01).
Figure 1. Adherence to diet and microbiome at baseline. A) Adherence to diet evaluated by the plasma metabolomics readout. B) Alpha diversity (Shannon) at baseline is significantly higher in patients with at least 50% pain improvement (R) compared to patients with a less than 50% improvement in pain (NR). C) Change in alpha diversity from day-14 (before diet) to day +14 (after diet) in R versus NR, assessed by at least 50% pain improvement. Boxes represent group means with standard deviations. D) Correlation (Spearman rank correlation) of bacterial genus with clinical outcomes at baseline (CDAI – clinical disease activity index, DAS28CRP – disease activity score using 28 joint count and C reactive protein). The significant correlations are marked with * (p < 0.05) or ** (p < 0.01).
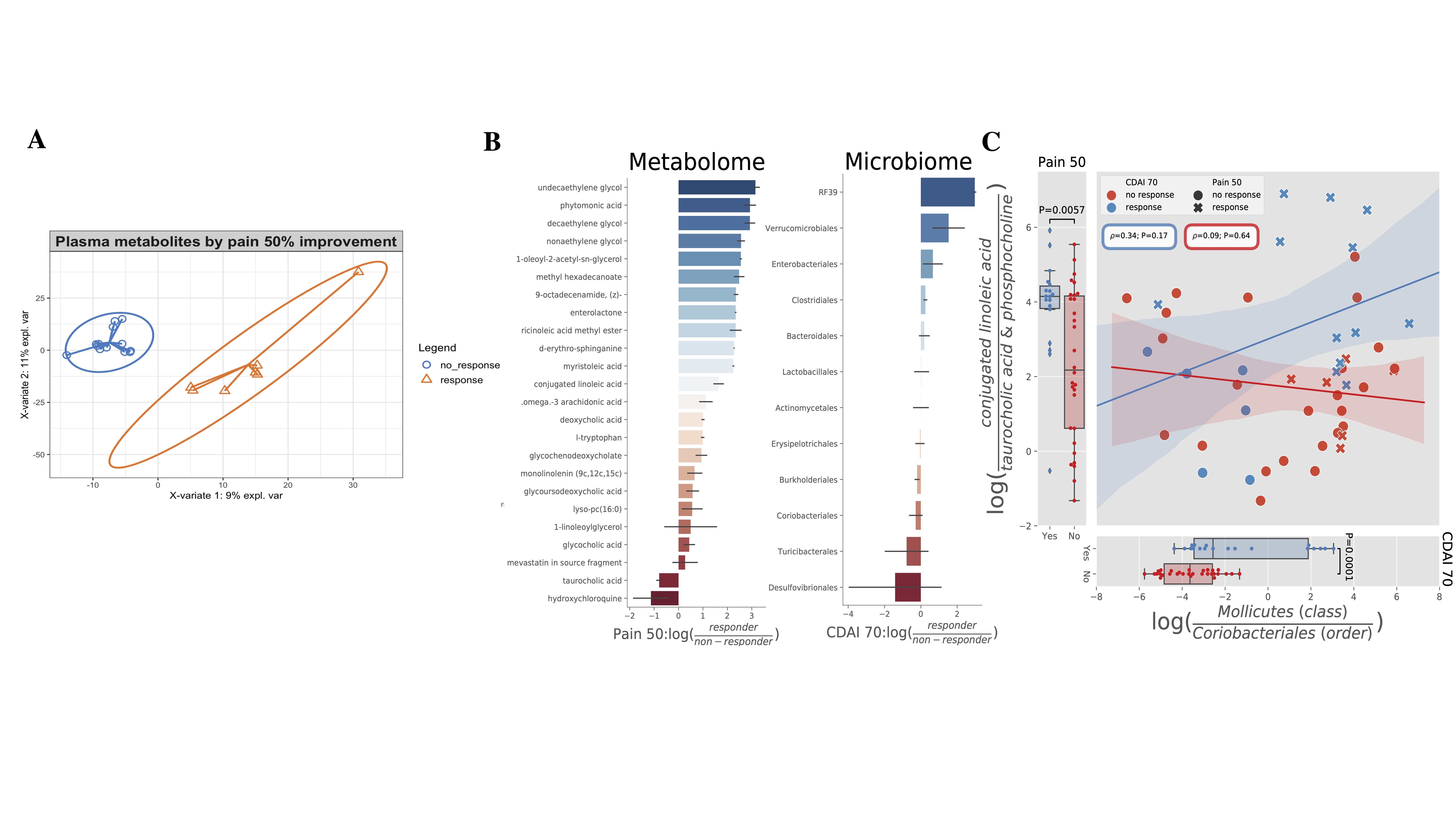 Figure 2. Plasma metabolome is different in R versus NR. A) Partial least square regression showing that plasma metabolome is different in R versus NR. B) Differential metabolites (left) and microbes (right) by chemical name and taxonomic order (y-axis) between responders and non-responders (x-axis). C) In the analysis of differential metabolomics Pain50 responders are enriched in conjugated linoleic acid (CLA) compared to non-responders (NR), while the Pain50 NR are enriched in the biliary metabolite taurocholic acid and phosphocholines compared to R (left). This trend was verified with a log-ratio between the two metabolite groups which is significant between R and NR (T=2.9, P=0.006). In the analysis of differential sub-operational taxonomic units by CDAI 70, Mollicutes were found to be enriched and Coriobacteriales depleted in R compared to NR (right). This trend was verified with a log-ratio between the two taxonomic orders which is significant between R and NR (T=4.22, P=0.0001). Error bars in all figures shaded by standard error. Correlation performed through spearman rank-order correlation. Log-ratio significance determined through two-sided T-test.
Figure 2. Plasma metabolome is different in R versus NR. A) Partial least square regression showing that plasma metabolome is different in R versus NR. B) Differential metabolites (left) and microbes (right) by chemical name and taxonomic order (y-axis) between responders and non-responders (x-axis). C) In the analysis of differential metabolomics Pain50 responders are enriched in conjugated linoleic acid (CLA) compared to non-responders (NR), while the Pain50 NR are enriched in the biliary metabolite taurocholic acid and phosphocholines compared to R (left). This trend was verified with a log-ratio between the two metabolite groups which is significant between R and NR (T=2.9, P=0.006). In the analysis of differential sub-operational taxonomic units by CDAI 70, Mollicutes were found to be enriched and Coriobacteriales depleted in R compared to NR (right). This trend was verified with a log-ratio between the two taxonomic orders which is significant between R and NR (T=4.22, P=0.0001). Error bars in all figures shaded by standard error. Correlation performed through spearman rank-order correlation. Log-ratio significance determined through two-sided T-test.
To cite this abstract in AMA style:
Coras R, Martino C, Gauglitz J, Tripathi A, Jarmusch A, Cedola F, Fernandez Bustamante M, Agustín-Perez M, Alharthi M, Lee S, Singh A, Choi S, Rivera T, Nguyen K, Shekhtman T, Holt T, Golshan S, Knight R, Dorrestein P, Guma M. Rheumatoid Arthritis Improvement After Exposure to an Anti-Inflammatory “ITIS” Diet Is Associated with Changes of Gut Microbiome and Systemic Metabolome [abstract]. Arthritis Rheumatol. 2020; 72 (suppl 10). https://acrabstracts.org/abstract/rheumatoid-arthritis-improvement-after-exposure-to-an-anti-inflammatory-itis-diet-is-associated-with-changes-of-gut-microbiome-and-systemic-metabolome/. Accessed .« Back to ACR Convergence 2020
ACR Meeting Abstracts - https://acrabstracts.org/abstract/rheumatoid-arthritis-improvement-after-exposure-to-an-anti-inflammatory-itis-diet-is-associated-with-changes-of-gut-microbiome-and-systemic-metabolome/
