Session Information
Session Type: Abstract Session
Session Time: 5:00PM-5:50PM
Background/Purpose: To utilize novel methodologies to explore subgroups within the OAI clinical data.
Methods: From the OAI baseline dataset (n=4796 individuals with or at risk of knee OA/9592 knees; 116 clinical and demographic features; https://nda.nih.gov/oai/; AllClinical00), we excluded uninformative variables and features/knees with missing data leaving 86 features from 3322 people (6461 knees) for analyses. Continuous variables were transformed using a procedure to remove skewness and standardized (Feng 2016; doi: 10.1002/sta4.104).
Biclustering (Cheng 2000; PMID 10977070) is particularly useful when different subsets of features are informative for some, but not all, knees. This method allows simultaneous clustering of the rows (knees) and columns (features) of the matrix, and automated discovery of similarities based on these attributes. A bicluster is then defined as a sub-matrix that is well-fit (i.e., larger R2) by two-way ANOVA, in that the entries are close to a common value with adjustments for the rows and columns. We chose the parameters of the algorithm to balance competing desires for biclusters: strongly similar knees and sufficient sample size to provide potentially useful phenotypes.
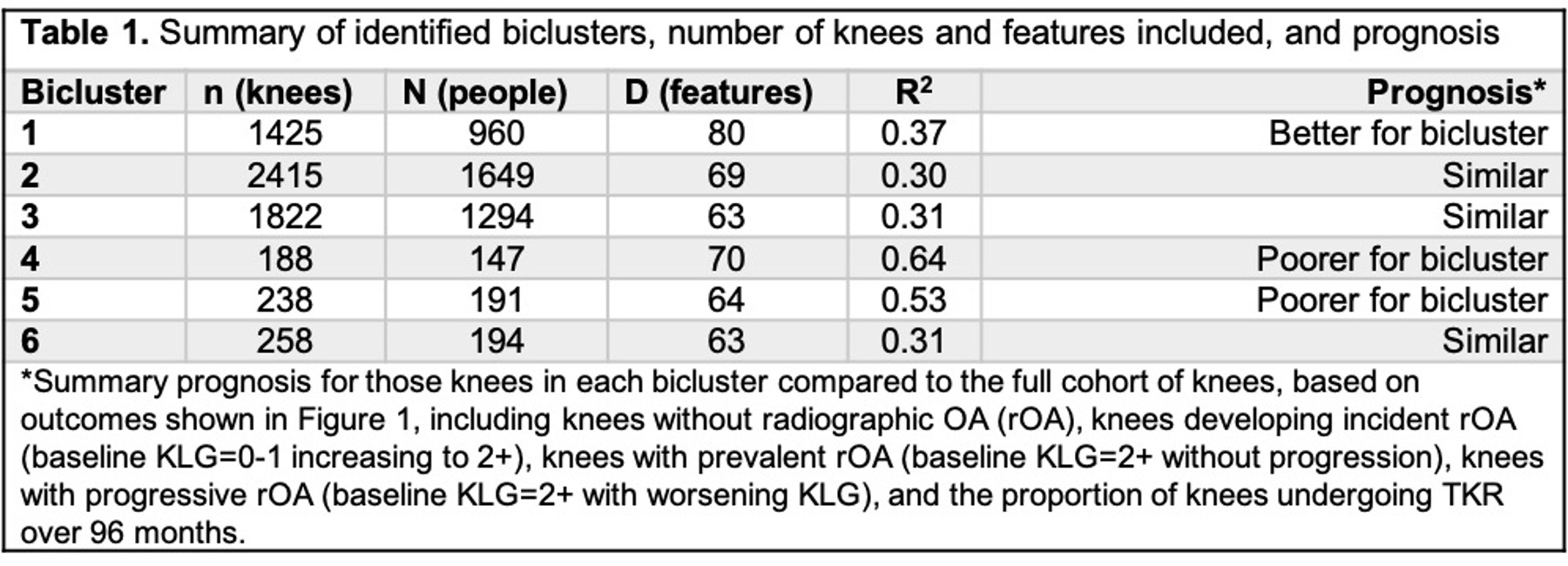
After identifying potential biclusters, we characterized them by comparing the knees within the bicluster to the overall population of knees based on marginal distributions of features as well as outcomes (defined in Table 1) such as prevalent, incident, and progressive knee OA and provision of knee replacement (TKR) from baseline to 96 months.
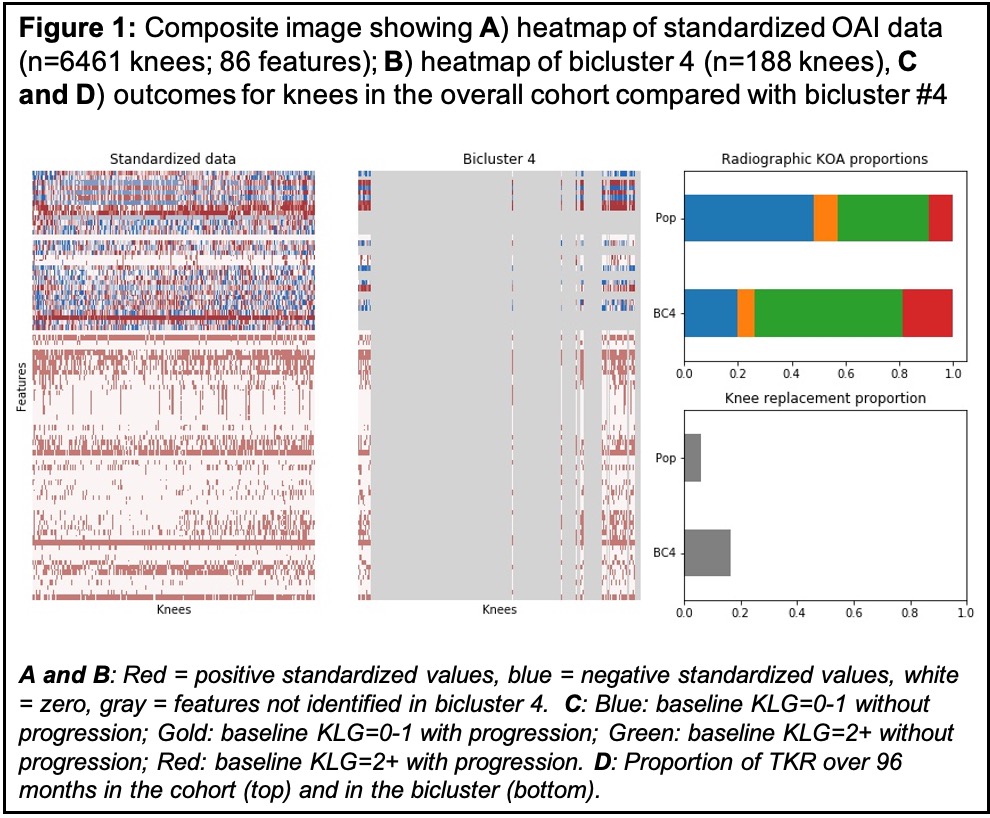
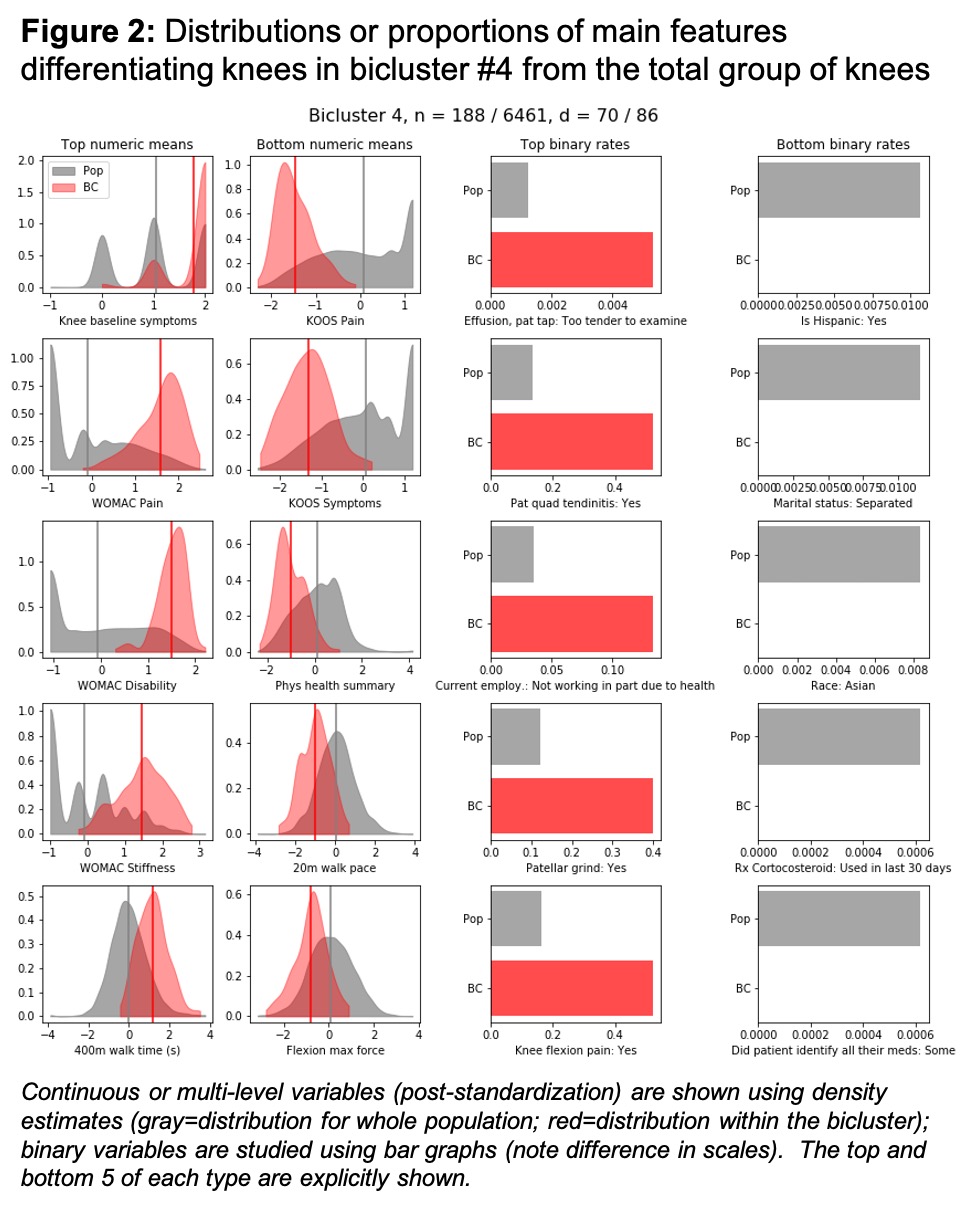
Results: We found 6 biclusters (chosen until the clusters became too small to be useful, Table 1). As an example, bicluster #4 contained 188 knees (from 147 people) and 70 features (Fig 1B) and was characterized by greater frequency of both progressive OA (Fig 1C) and TKR (Fig 1D). We then compared the distribution of baseline features between the knees in bicluster #4 and the total population of knees (Fig 2). Features that were higher among the knees in bicluster #4 versus the overall population of knees included knee symptoms, WOMAC scores, 400-meter walk time, knee tenderness on exam, patellar quadriceps tendinitis, inability to work for health reasons, presence of patellar grind, and knee flexion pain. In contrast, knees included in the bicluster had lower values for KOOS (to be expected as the coding is opposite that of WOMAC), SF-12 physical summary scale, 20-meter walk speed, and maximal isometric flexion strength. These knees were less likely to be from Hispanic or Asian participants, individuals who were separated (versus any other marital status), and less frequently reported IA corticosteroids in the last 30 days.
Conclusion: We identified several biclusters (groups of features and knees) within the baseline OAI data with varying prognoses. Such biclusters may represent potential phenotypes (e.g., progressor phenotype(s)) within the larger cohort. Application of novel methodologies can provide new insights into OA phenotypes and development or progression of disease.
To cite this abstract in AMA style:
Nelson A, Keefe T, Schwartz T, Loeser R, Golightly Y, Arbeeva L, Marron J. Biclustering Reveals Potential Knee Osteoarthritis Phenotypes in Exploratory Analyses: Data from the Osteoarthritis Initiative [abstract]. Arthritis Rheumatol. 2020; 72 (suppl 10). https://acrabstracts.org/abstract/biclustering-reveals-potential-knee-osteoarthritis-phenotypes-in-exploratory-analyses-data-from-the-osteoarthritis-initiative/. Accessed .« Back to ACR Convergence 2020
ACR Meeting Abstracts - https://acrabstracts.org/abstract/biclustering-reveals-potential-knee-osteoarthritis-phenotypes-in-exploratory-analyses-data-from-the-osteoarthritis-initiative/