Session Information
Date: Sunday, November 7, 2021
Title: Plenary II (0956–0961)
Session Type: Plenary Session
Session Time: 11:45AM-12:00PM
Background/Purpose: Multi-system Inflammatory Syndrome in Children (MIS-C) is a major complication of the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) pandemic in pediatric patients. Children with MIS-C present with varying degrees of cardiovascular and hyperinflammatory symptoms. We performed a comprehensive analysis of the plasma proteome of more than 1400 proteins in children with SARS-CoV-2. We hypothesized that the proteome would reflect heterogeneity in hyperinflammation and vascular injury, and further identify pathogenic mediators of disease.
Methods: Between April 2020 and October 2020, we enrolled 63 hospitalized patients with MIS-C (N=22), Severe COVID-19 (“Severe”, N=15)) or asymptomatic or mild SARS-CoV-2 infection (“Minimal”, N=26). In addition, we included remnant plasma samples from otherwise healthy patients (“Healthy”, N=25). We utilized the Olink Explore 1536/384 protein biomarker platform to interrogate the plasma proteome of all 88 patients. All statistical analyses were performed in RStudio (RStudio, PBC, Boston, MA).57 Data were assumed to be non-parametric unless normality was demonstrated.
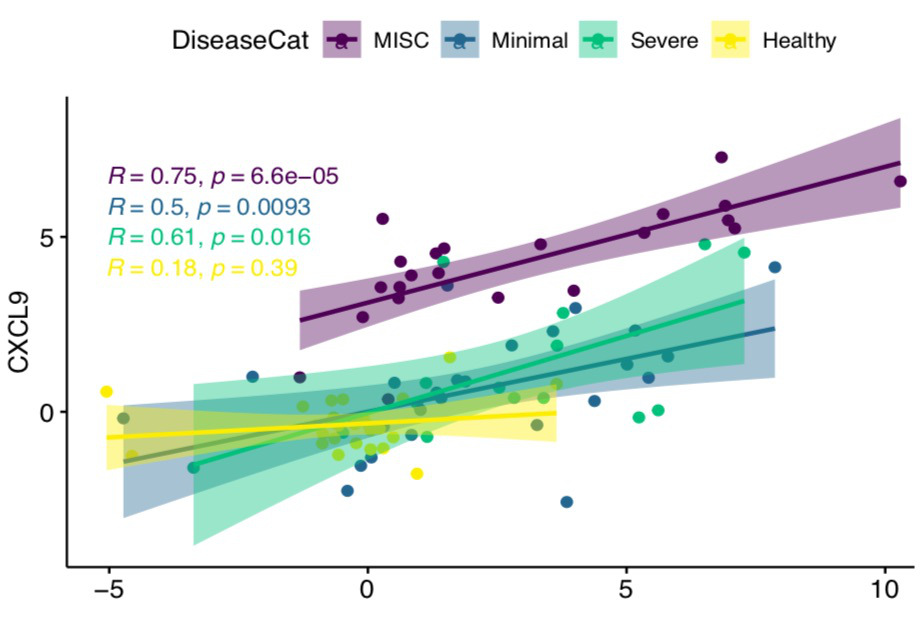
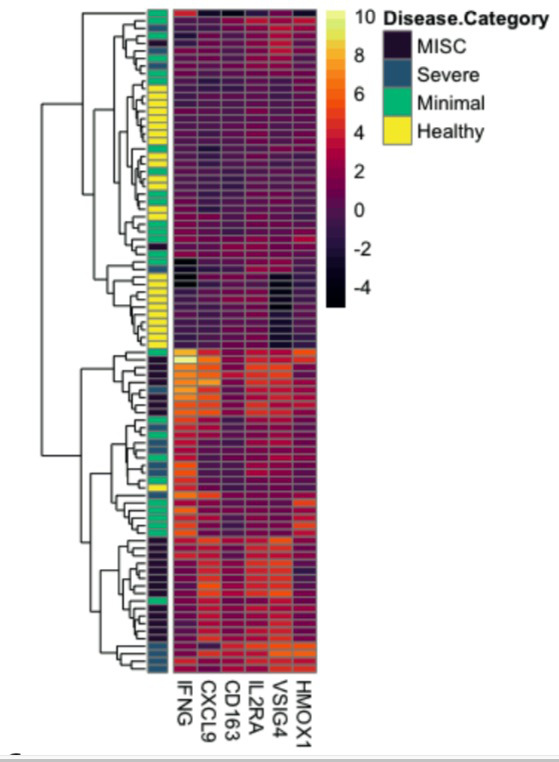
Results: To understand how the overall proteome of patients with MIS-C changes over time, we first created a PCA mapping all proteins and all patients. We then PCA transformed data for convalescent samples on that space. Convalescent samples shifted towards the healthy controls, implying that following treatment, the proteome in MIS-C patients returns towards a baseline state. We examined CXCL9 as the key protein associated with IFNg response and found that all patients with SARS-CoV-2 infection showed a positive CXCL9 response to increasing IFNg. Strikingly, MIS-C patients had a disproportionately high CXCL9 response to IFNg compared to the other groups. Consistent with a derepression of IFNg signaling, MIS-C patients expressed less TRIM21 compared to healthy controls. Unsupervised hierarchical clustering revealed two distinct groups of MIS-C patients that had elevations in multiple Macrophage Activation Syndrome (MAS) markers. These two clusters primarily differed by their IFNg expression with an IFNg-high versus IFNg-low group evident. Hierarchical clustering identified that SC5B9 correlated most highly with PLA2G2A, PDGFC, SELE, CALCA, NOS3, VWA1 and TYMP, demonstrating a signature consistent with thrombotic microangiopathy (TMA). Patients with elevations in MAS or TMA markers were more likely to meet clinical criteria for MAS or TMA respectively. Interestingly, MISC patients with an Interferon gamma low signature were at higher risk of requiring inotropes.
Conclusion: We used proteomic analysis to begin to unravel the complex and heterogeneous pathophysiology associated with MIS-C. We identify PLA2G2A as a candidate biomarker for MIS-C and show that PLA2G2A is associated with clinical features of TMA. We demonstrate that MIS-C patients are characterized by a disproportionately high CXCL9 response to IFNg, implying a dysregulated response to IFNg. Patients with MIS-C showed heterogeneity based on IFNg expression, and surprisingly, patients with higher IFNg levels were less likely to require inotropes.
To cite this abstract in AMA style:
Diorio C, Shraim R, Vella L, Giles J, Baxter A, Oldridge D, Canna S, Henrickson S, Mcnerney K, Balamuth F, Burudpakdee C, Lee J, Leng T, Farrell A, Lambert M, Sullivan K, Wherry J, Teachey D, Bassiri H, Behrens E. Proteomic Profiling of MIS-C Patients Reveals Heterogeneity Relating to Interferon Gamma Dysregulation and Vascular Endothelial Dysfunction [abstract]. Arthritis Rheumatol. 2021; 73 (suppl 9). https://acrabstracts.org/abstract/proteomic-profiling-of-mis-c-patients-reveals-heterogeneity-relating-to-interferon-gamma-dysregulation-and-vascular-endothelial-dysfunction/. Accessed .« Back to ACR Convergence 2021
ACR Meeting Abstracts - https://acrabstracts.org/abstract/proteomic-profiling-of-mis-c-patients-reveals-heterogeneity-relating-to-interferon-gamma-dysregulation-and-vascular-endothelial-dysfunction/