Session Information
Date: Saturday, November 6, 2021
Title: SLE – Diagnosis, Manifestations, & Outcomes Poster I: Diagnosis (0323–0356)
Session Type: Poster Session A
Session Time: 8:30AM-10:30AM
Background/Purpose: Around 40-70% of patients with systemic lupus erythematosus (SLE) develop lupus nephritis (LN). LN is often underdiagnosed and early diagnosis is crucial for reducing kidney damage. Various tests can diagnose LN including kidney biopsy, but LN symptoms can vary greatly between patients making diagnosis challenging especially in the early stages. We aimed to develop a machine learning (ML)-based approach to classify SLE patients at high risk of LN to assist rheumatologists in their diagnosis.
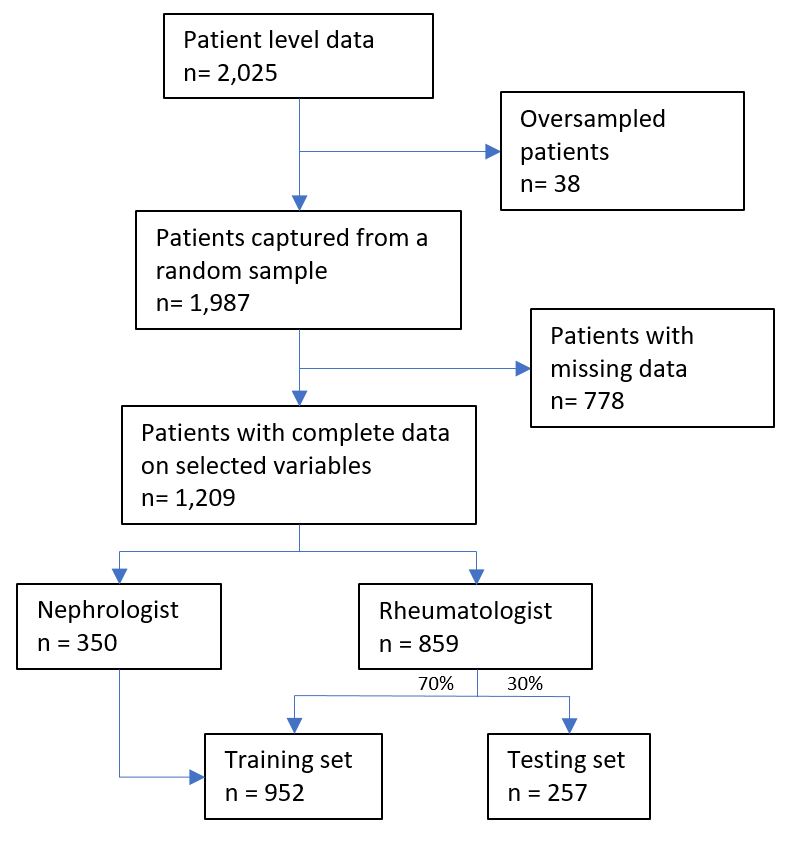
Methods: Data were drawn from the Adelphi Lupus Disease Specific programme™, a point-in-time survey of rheumatologists and nephrologists and their SLE/LN patients conducted in France, Germany, Italy, Spain, the United Kingdom and the United States of America from Aug–Oct 2015. Rheumatologists and nephrologists each completed record forms for their next 5 consecutive adult SLE and LN patients, respectively, capturing demographic and clinical characteristics. Patients were categorized as LN or non-LN based on physicians’ assessment.
We subset data for training and testing the ML model, selecting 354 variables (from a total of 30 record form questions) based on specialist knowledge of LN pathophysiology. To ensure maximum discriminative power, the training set comprised all nephrologist-diagnosed patient and 70% rheumatologist-diagnosed patient data; the testing set comprised the remaining 30% rheumatologist-diagnosed patient data. Training and hyperparameter tuning were done using 10-fold cross-validation with R and Tidymodels Package. Model predictive performance was assessed by receiver operating characteristic area under the curve (AUC) metric.
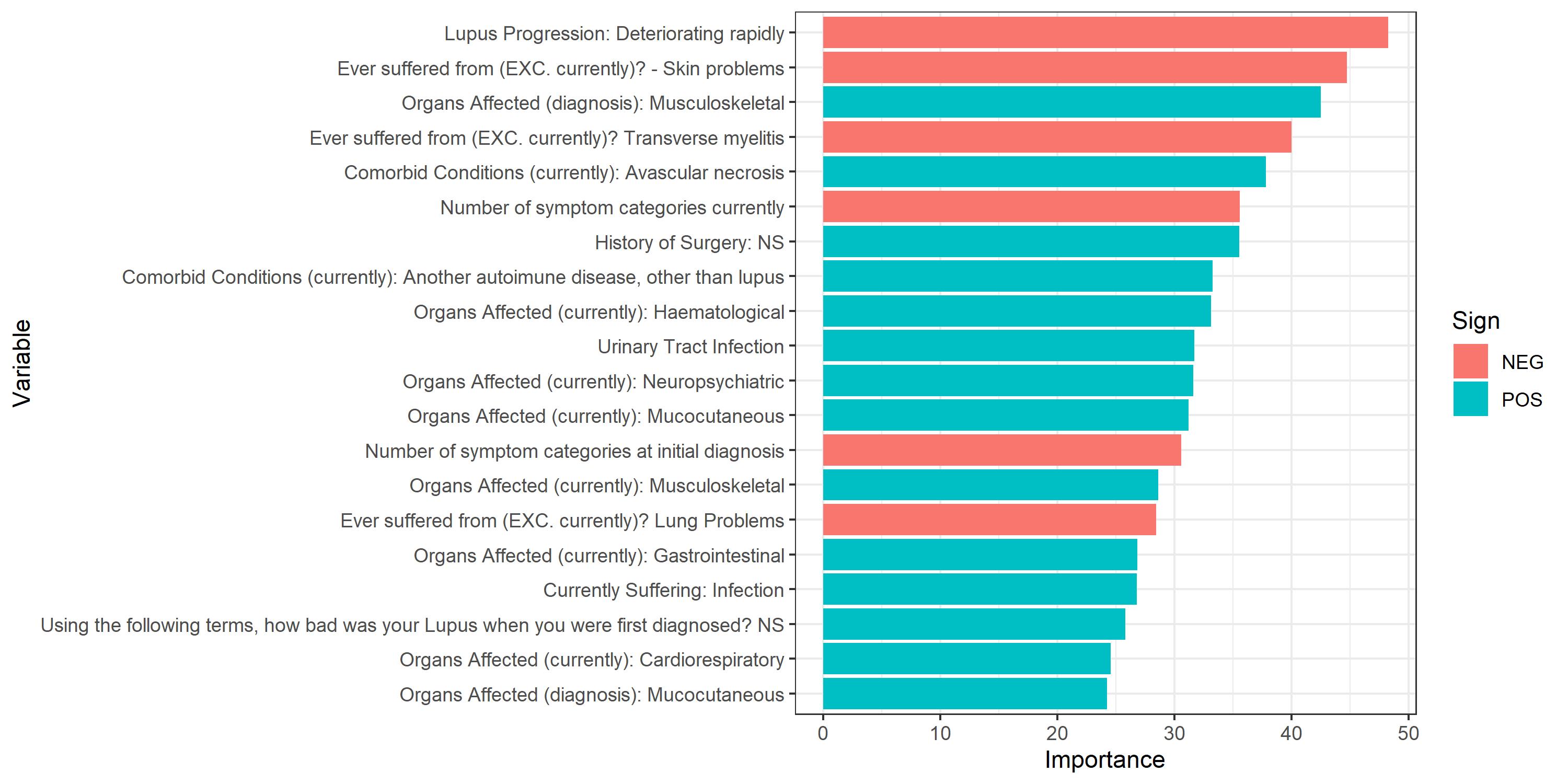
Results: The model used data collected by 304 physicians on 1,209 SLE patients including both LN (n=483) and non-LN (n=726) patients (Fig. 1). After testing 2,520 models using the 6 most commonly used classification algorithms we found that the penalized logistic regression (PLR) with elastic net regularization predicted which SLE patients were at highest risk of LN (AUC: 0.91, with negative/positive predictive values of 0.96 and 0.68, respectively) in the testing set. The 3 variables that best predicted LN risk were disease progression, skin problems (both contributing negatively to risk) and musculoskeletal organs affected at diagnosis (with a positive contribution to risk, Fig. 2).
Conclusion: PLR successfully predicted SLE patients’ LN risk using real-world data. Although linear discriminant analysis had a slightly better AUC (0.92), PLR results are easier to interpret, making it a suitable model to support rheumatologists identifying LN patients within the SLE population to guide further testing and treatment.
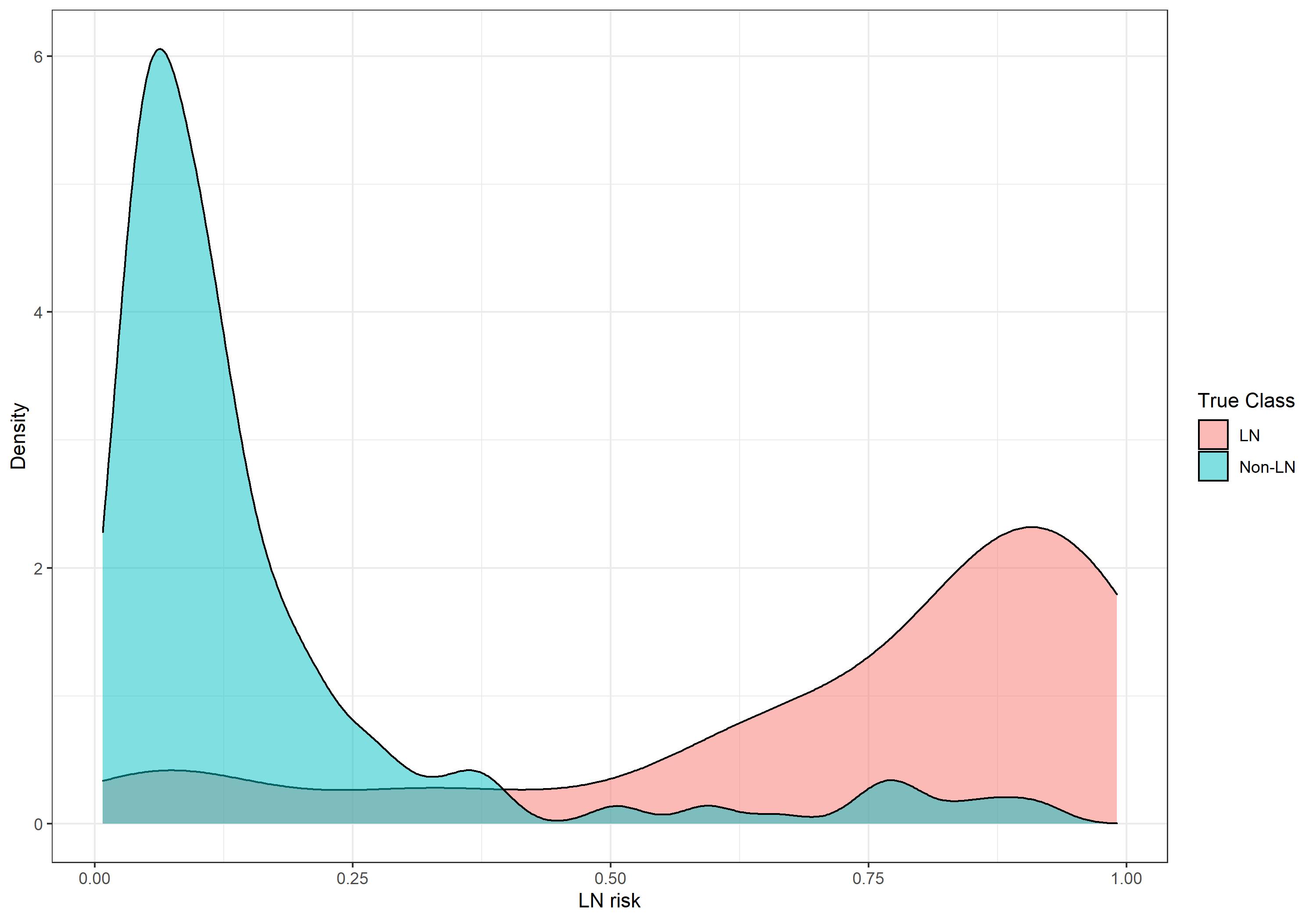
The PLR model had better discriminative power in non-LN vs LN patients. This may have resulted from undiagnosed LN patients remaining within the non-LN patient pool and thus generating false positives, since these high LN risk non-LN patients shared a number of characteristics with LN patients (Fig. 3).
Predictive analysis has shown promising results for LN risk assessment in SLE patients using real-world data, we recommend further prospective validation is required prior to use in a clinical setting.
Variables in red contributed negatively to lupus nephritis risk prediction
NEG, negative; POS, positive
LN, lupus nephritis
To cite this abstract in AMA style:
Teixeira B, Bell D, Holbrook T, Hoskin B. Machine Learning: Identifying Lupus Nephritis Within Systemic Lupus Erythematosus in the Real World [abstract]. Arthritis Rheumatol. 2021; 73 (suppl 9). https://acrabstracts.org/abstract/machine-learning-identifying-lupus-nephritis-within-systemic-lupus-erythematosus-in-the-real-world/. Accessed .« Back to ACR Convergence 2021
ACR Meeting Abstracts - https://acrabstracts.org/abstract/machine-learning-identifying-lupus-nephritis-within-systemic-lupus-erythematosus-in-the-real-world/