Session Information
Session Type: Poster Session (Sunday)
Session Time: 9:00AM-11:00AM
Background/Purpose: Filgotinib (FIL), an oral, selective, Janus Kinase 1 (JAK1) inhibitor was effective in Phase 3 studies of active RA in patients (pts) with inadequate response or intolerance to biologic DMARDs (bDMARD-IR; FINCH-2 ClinicalTrials.gov Identifier: NCT02873936). A proteomic and transcriptomic study was conducted to elucidate the effect of FIL on molecular mechanisms and the underlying pathobiology of RA.
Methods: Whole blood was collected from FINCH-2 RA pts on either a stable dose of placebo + MTX (PBO; n = 148), or once daily FIL 100 mg (n = 153), or 200 mg (n = 148) + MTX at baseline, week 4 and week 12. Samples from demographically matched healthy volunteers (HV; n = 50) were also evaluated. Serum concentrations of 28 cytokines known to be associated with RA pathology were quantified by ELISA-based assays. Globin depleted whole-blood mRNA was sequenced using Illumina HiSeq 2500. Differential gene expression analysis was applied comparing baseline RA pts data to HVs and interpreted into pathways using Gene Set Enrichment Analysis (GSEA). Further, we calculated gene signature (GS) scores using single sample GSEA (ssGSEA) to evaluate a reversion to healthy for Hallmark gene sets that had previously shown the strongest magnitude of FIL-associated effects (Taylor et al. EULAR 2019). Each GS was tested (Wilcoxon rank sum) comparing treatment arms with time-paired subjects against HVs. All cytokines and GSs were standardized to HV values. Cytokines and GSs not statistically different between RA pts after treatment and HVs were considered to have “reversed” toward a HV phenotype.
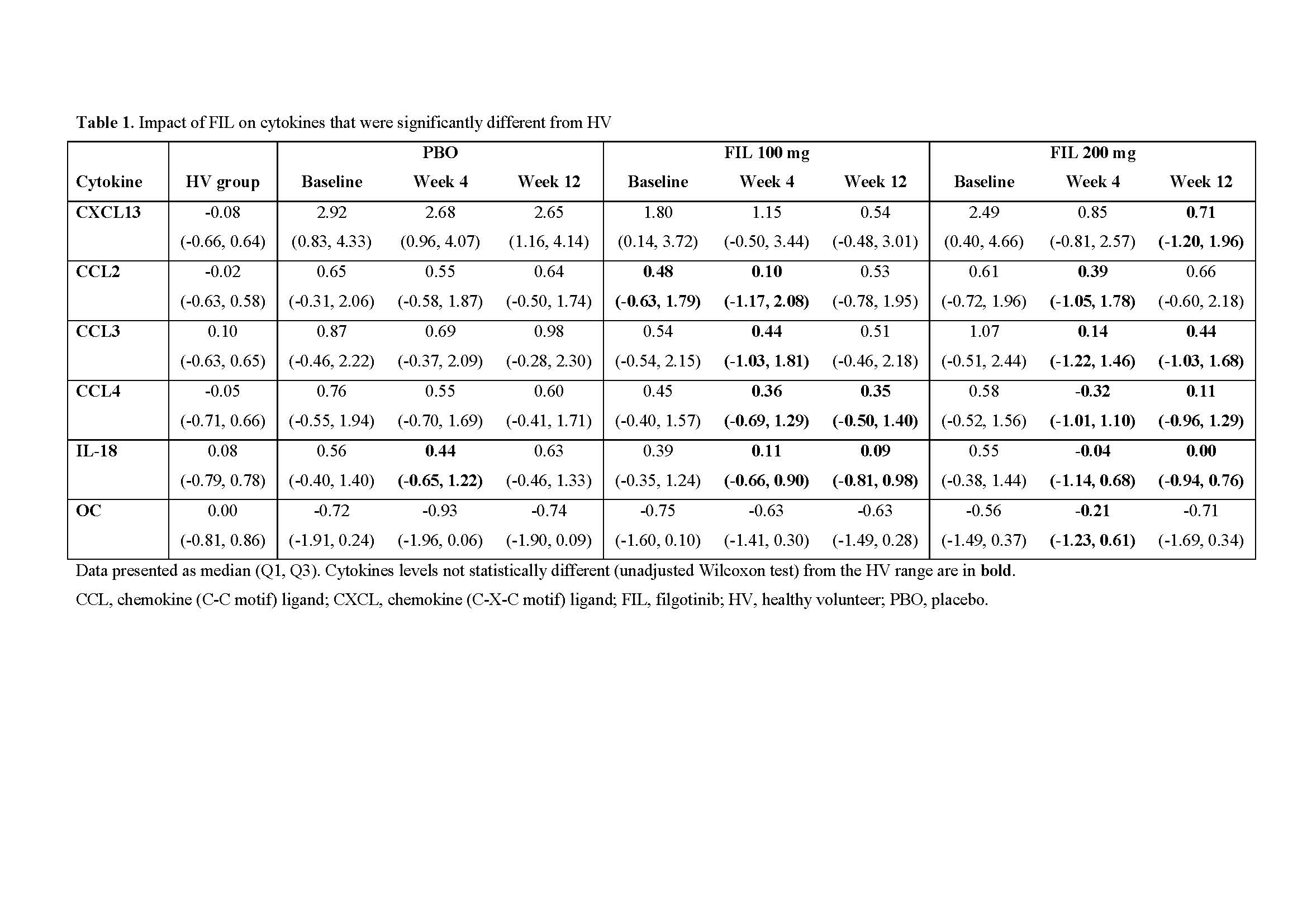
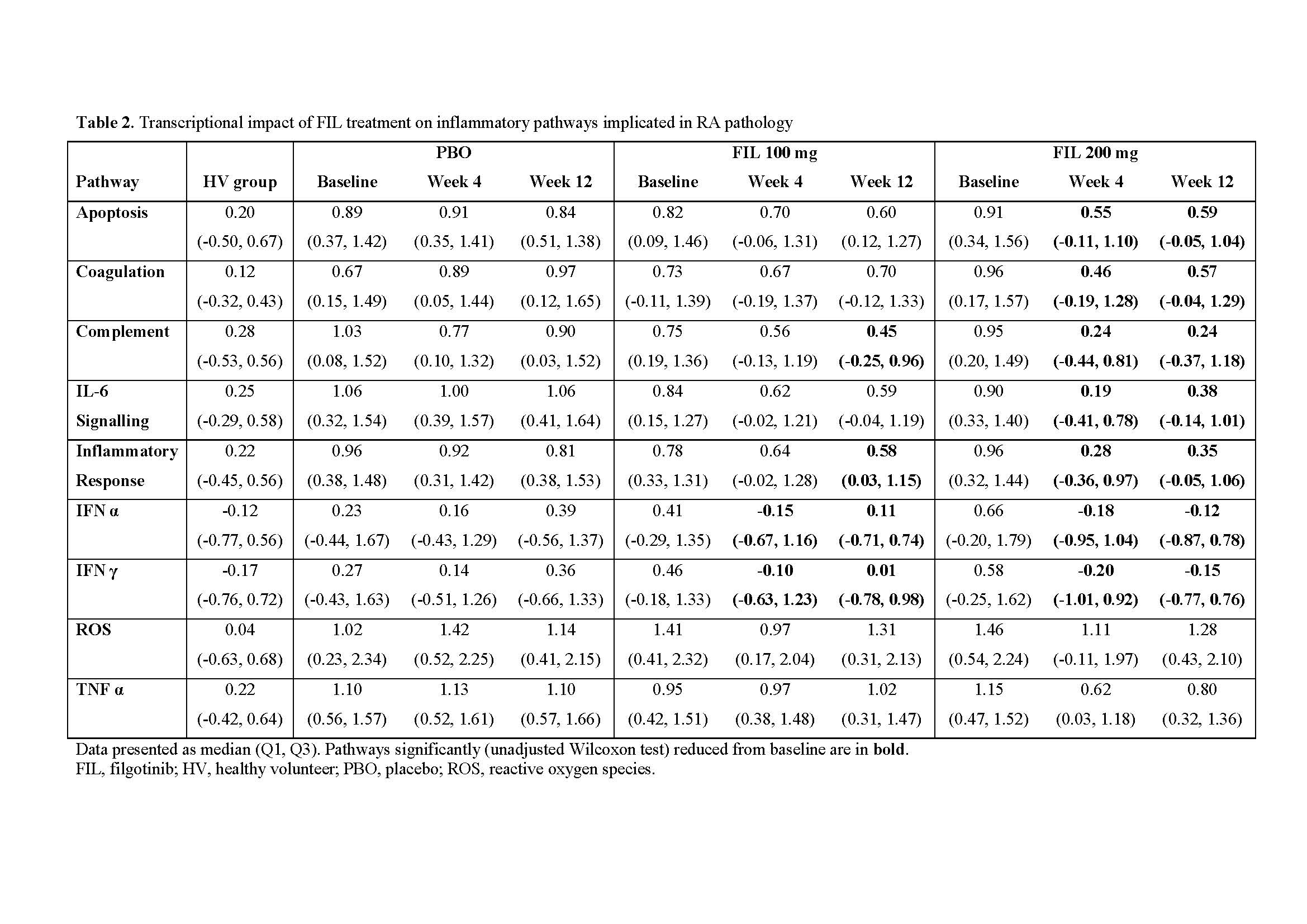
Results: Among 28 cytokines evaluated in baseline RA serum, 18 were significantly different from HV. Of these, 6 (chemokine (C-X-C motif) ligand [CXCL13], chemokine (C-C motif) ligand 2 [CCL2], CCL3, CCL4, IL-18, and Osteocalcin) reversed towards a HV phenotype after FIL treatment (Table 1). In contrast, only IL-18 reached HV levels in PBO treated pts (week 4). Of 15,189 genes with expression data, 3,532 (23%) were significantly increased and 4,408 (29%) significantly decreased in pts with RA relative to HV (false discovery rate < 0.05). The baseline transcriptional profile of RA pts was enriched with pathways implicated in RA pathology, including inflammatory responses and IFN, IL-6, TNFα signalling. Reversal of transcriptional profiles was broadly dose- and time-dependent, with fewer genes differentially expressed from HVs after 12 weeks of FIL compared with those receiving PBO. Expression of IFN response genes in pts who received FIL was statistically indistinguishable from HVs after 12 weeks. Interestingly, reactive oxygen species and TNFα signalling did not show this trend of reversal towards HV.
Conclusion: Differences in the peripheral molecular profile were observed between bDMARD-experienced RA pts and HV. While an overall trend toward the non-diseased molecular profile was observed following FIL, only a subset of cytokines and pathways were statistically indistinguishable from HV after 12 weeks. These results support the clinical efficacy of FIL within the bDMARD-experienced population and provide evidence for the reversal of disease activity to a healthier molecular profile in the periphery.
To cite this abstract in AMA style:
Taylor P, Elboudwarej E, Downie B, Vestergaard L, Liu J, Mirza A, Hawtin R. bDMARD-experienced Filgotinib-treated Patient Samples Exhibit a Partial Reversion to the Peripheral Molecular Profile of a Demographically Matched Healthy Population [abstract]. Arthritis Rheumatol. 2019; 71 (suppl 10). https://acrabstracts.org/abstract/bdmard-experienced-filgotinib-treated-patient-samples-exhibit-a-partial-reversion-to-the-peripheral-molecular-profile-of-a-demographically-matched-healthy-population/. Accessed .« Back to 2019 ACR/ARP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/bdmard-experienced-filgotinib-treated-patient-samples-exhibit-a-partial-reversion-to-the-peripheral-molecular-profile-of-a-demographically-matched-healthy-population/