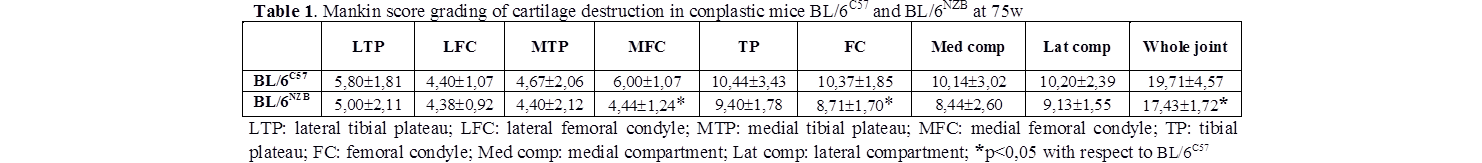Session Information
Session Type: ACR Poster Session B
Session Time: 9:00AM-11:00AM
Background/Purpose:
Different studies showed interesting genetic associations between mtDNA haplogroups and OA-related features, including prevalence, incidence and progression of the disease. The aim of this work is to study the influence of mtDNA variation in the degree of joint deterioration of knees from a conplastic mouse model of aging
Methods:
For this purpose, mtDNAs from C57BL/6 and NZB/OlaHsd mice were used. These mtDNAs differ by 12 missense mutations, 10 non-coding-region mutations, 8 ribosomal RNA mutations and 4 transfer RNA mutations. A conplastic mice strain was developed with the C57BL/6 nuclear genome and the NZB/OlaHsd mtDNA (named BL/6NZB) to compare with the original C57BL/6 strain (named BL/6C57) in animals of 25 and 75 weeks (25w and 75w). A total of 38 limbs from 19 mice were processed to perform different histologic analysis: 8 BL/6NZB 25w, 10 BL/6NZB 75w, 10 BL/6C57 25w and 10 BL/6C57 75w. Appropriate non-parametric statistical procedures were applied using SPSS v24
Results:
Cartilage and epiphyseal plate were the joint tissues with the most relevant findings. Mankin score data showed significantly increased values in all knees from both animal models at 75 weeks compared with 25 weeks (p<0,001), confirming the aging of the joint. When 75-week mice were selected (table 1), the BL/6C57 strain showed a significantly increased score in whole joint (p=0.038), medial femoral condyle (p=0,015) and femoral condyle (p=0,021) than BL/6NZB strain. Safranin-O/Fast-green ratio value at 75w was higher in the medial compartment (both tibial plateau and femoral condyle) of BL/6NZB compared with BL/6C57; however, only the differences detected in the medial compartment of the tibial plateau reached the statistical significance (p<0,001), whilst the differences detected in the femoral condyle borderline the statistical significance (p=0,091).
The width of the epiphyseal plate was analyzed in both femur and tibia bones. The results showed significantly decreased values in BL/6C57 75w compared with the same strain at 25 weeks in femoral condyle (p=0,049) and tibial plateau (p<0,001); however, these differences were not observed in animals belonging to BL/6NZB strain. In addition, the BL/6C57 75w strain also showed significantly lower values in tibial plateau than BL/6NZB strain at the same age (p=0,032).
Conclusion:
mtDNA variation plays an important role in the process of joint deterioration associated to age, leading to consider both mitochondria and mtDNA as potential therapeutic targets and complementary biomarker, respectively, for the age-associated phenotype of OA
To cite this abstract in AMA style:
Rego-Pérez I, Lechuga-Vieco AV, Scotece M, Filgueira-Fernández P, Pertega S, Enríquez JA, Blanco FJ. A Conplastic Mice Model of Aging Reveals That Mitochondrial DNA Variation Influences the Process of Joint Deterioration [abstract]. Arthritis Rheumatol. 2018; 70 (suppl 9). https://acrabstracts.org/abstract/a-conplastic-mice-model-of-aging-reveals-that-mitochondrial-dna-variation-influences-the-process-of-joint-deterioration/. Accessed .« Back to 2018 ACR/ARHP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/a-conplastic-mice-model-of-aging-reveals-that-mitochondrial-dna-variation-influences-the-process-of-joint-deterioration/