Session Information
Session Type: Poster Session (Tuesday)
Session Time: 9:00AM-11:00AM
Background/Purpose: Psoriasis is an inflammatory skin disease. Psoriatic arthritis (PsA) is an inflammatory arthritis that affects 30% of psoriasis patients. The heterogeneity of psoriatic disease (PsD- psoriasis and PsA) is partly explained by genetic heterogeneity: human leukocyte antigen (HLA)-C*06 is the strongest genetic risk factor for psoriasis without arthritis (PsC), while HLA-B*27 and other HLA-B alleles are more strongly associated with PsA. Yet, PsD pathogenesis is not fully understood, inspiring an interest in the role of an important environmental factor: the microbiome. We aimed to characterize the cutaneous microbiome of psoriatic plaques and compare its diversity between PsC and PsA patients and between the major PsD-associated HLA alleles.
Methods: Skin swabs were obtained from a psoriatic lesion on the extensor aspect of the elbow or knee, and from unaffected skin on the contralateral joint in PsC patients (psoriasis diagnosed by a dermatologist, evaluated by a rheumatologist to exclude PsA) and PsA patients satisfying CASPAR criteria. Control samples were taken from matched sites in healthy subjects. PsA patients were evaluated by a rheumatologist using a standard protocol to determine the number of affected joints, peripheral and/or axial joint involvement. Subsequently, microbial DNA was isolated from the swab samples, and high-throughput 16S ribosomal DNA sequencing was utilized to characterize the cutaneous microbiota. Molecular HLA Class I typing was done from DNA obtained from peripheral blood, and subjects were classified as being HLA-C*06 positive, HLA-B*27 positive or HLA-C*06 and -B*27 negative (termed double negative). All study subjects were Caucasian. Dada2 pipeline was used in the bioinformatic analysis to infer unique amplicon sequences from raw reads and remove chimeric sequences. QIIME v.1.9.1 was used to assign taxonomy with reference to an internally-designed 16S database, and to compute alpha-diversity. Biostatistical analysis was performed in R.
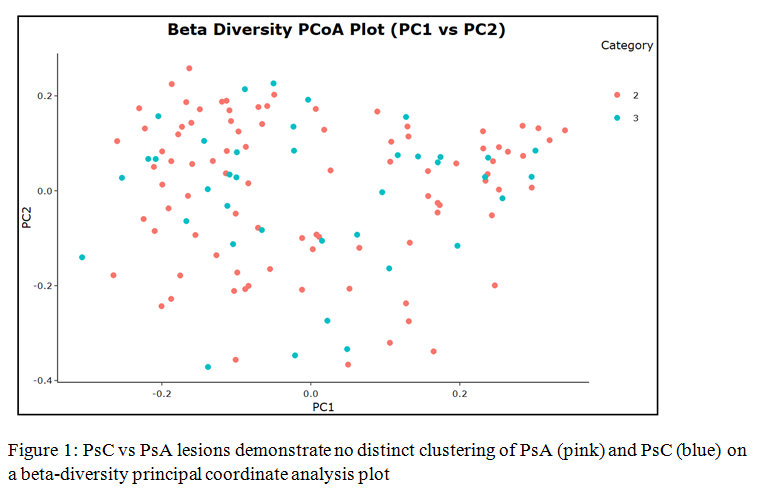
Results: We recruited 34 healthy controls (59% male), 55 PsC patients (47% male), and 136 PsA patients (63% male). 38 PsC and 90 PsA patients were included in this analysis after excluding subjects with recent use of topical ointments (≤7 days), antibiotics (≤3 months), and other related treatments. Linear model demonstrated that alpha diversity was significantly higher in lesional compared to unaffected samples in all PsD patients, as measured by Shannon diversity index (p=0.0106). However, there was no significant difference in lesional diversity between PsA and PsC patients (p=0.2244) (Figure 1). Interestingly, while there was no significant difference in alpha diversity between lesional swabs from HLA-B*27-positive compared to double negative patients (p=0.1401), it was shown to be significantly higher in lesional swabs from HLA-C*06-positive patients compared with both HLA-B*27-positive patients (p=0.0066) and double negative patients (p=0.0331) (Figure 2) by analysis of variance with Tukey’s correction for multiple comparisons.
Conclusion: The cutaneous lesional microbial diversity of patients with psoriatic disease may be influenced by the HLA-C*06 but not –B*27 alleles.
To cite this abstract in AMA style:
Yerushalmi M, Espin-Garcia O, Ye J, Pollock R, Rahmati S, Eder L, Silverberg M, Chandran V. The Cutaneous Microbiome of Psoriatic Disease Is Influenced by Disease Susceptibility HLA Alleles but Not Clinical Phenotype [abstract]. Arthritis Rheumatol. 2019; 71 (suppl 10). https://acrabstracts.org/abstract/the-cutaneous-microbiome-of-psoriatic-disease-is-influenced-by-disease-susceptibility-hla-alleles-but-not-clinical-phenotype/. Accessed .« Back to 2019 ACR/ARP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/the-cutaneous-microbiome-of-psoriatic-disease-is-influenced-by-disease-susceptibility-hla-alleles-but-not-clinical-phenotype/