Session Information
Date: Wednesday, November 13, 2019
Title: 6W023: Systemic Sclerosis & Related Disorder – Clinical III: Predictors of Outcome (2912–2917)
Session Type: ACR Abstract Session
Session Time: 11:00AM-12:30PM
Background/Purpose: We analyzed a phase 2 study designed to assess the efficacy of abatacept in patients with diffuse Systemic Sclerosis (SSc). In this work, we analyze data from ASSET and seek to confirm the hypothesis that patients in the inflammatory subset on abatacept show a significant decline in modified Rodnan Skin Score (mRSS), which is correlated to modulation of pathways related to the mechanism of action of abatacept.
Methods: SSc patients, who met 2013 ACR/EULAR criteria, were randomized to receive abatacept or placebo for 12 months. RNA sequencing was performed on 84 SSc patients at baseline, 3-month, and 6-month timepoints. Samples were assigned to an intrinsic gene expression subset (inflammatory, fibroproliferative, limited, or normal-like) using a Support Vector Machine (SVM) classifier. Treatment differences in longitudinal outcomes were assessed using linear mixed effect models. Improvement was defined as a 5 point or >20% change in mRSS between baseline and 12 months. A machine-learning approach, trained on gene-gene relationships in the skin, was used to identify features beyond those simply differentially expressed. Gene Set Enrichment Analysis (GSEA) was used to identify pathways enriched in comparison groups.
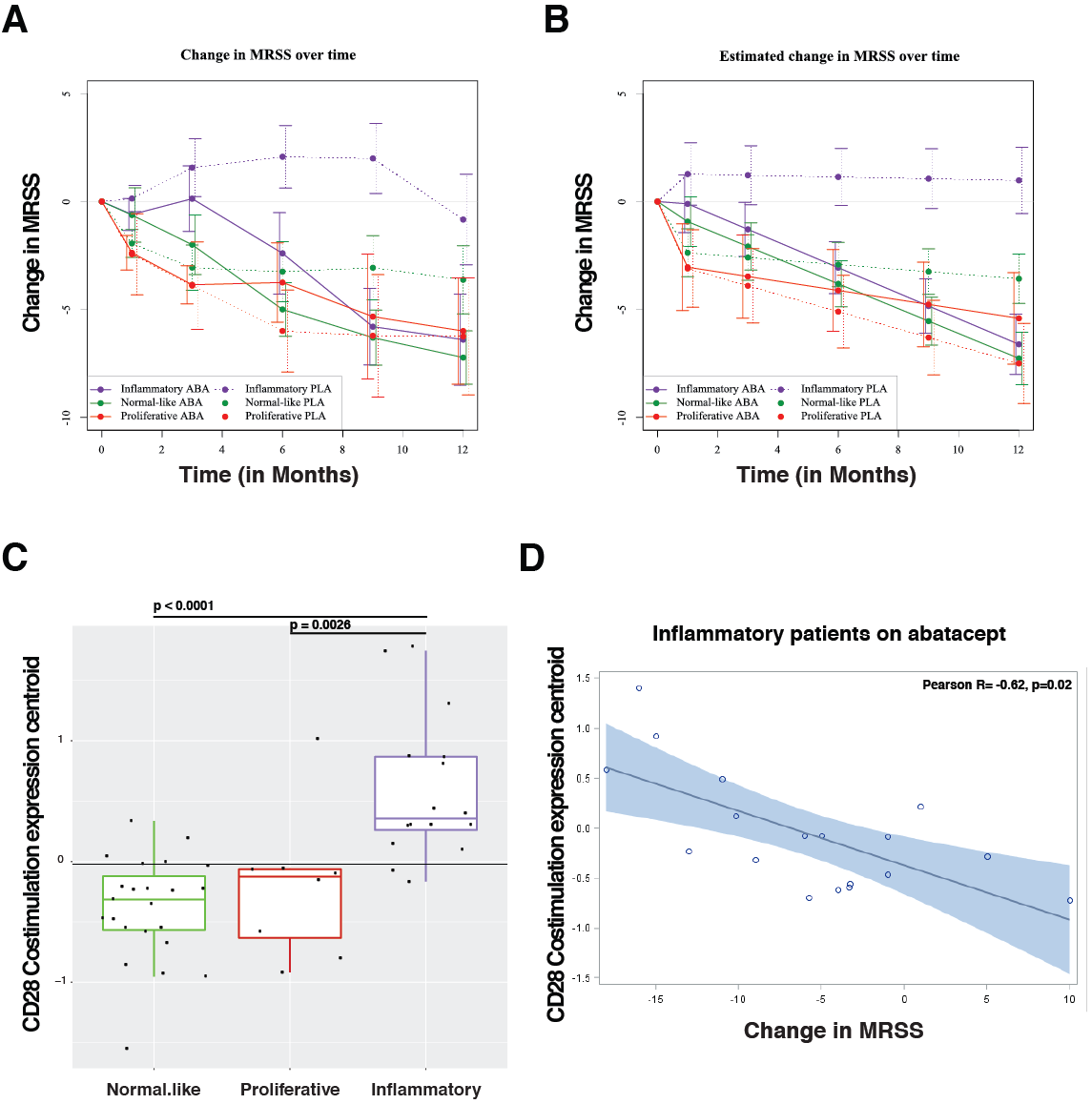
Results: Patients were assigned to intrinsic subset at baseline (33 inflammatory, 18 fibroproliferative, and 33 normal-like). Differences in trajectory for intrinsic subsets are evident (Figures 1a, 1b). In the abatacept arm, change in mRSS was most pronounced for the inflammatory (p< 0.001) and normal-like (p=0.03) subsets when compared to placebo. The gene-gene network from skin, returned CD86 as being highly related to genes that decrease in abatacept improvers. For gene expression analyses, the dataset was parsed for quality and patients that completed the trial, the remaining (n=140) biospecimens were analyzed. The pathway Costimulation by the CD28family decreases (FDR=5.88×10 -4 ) in patients that improve on abatacept; this decrease is specific to the inflammatory subset (FDR=0%) of patients. At baseline, patients in the inflammatory subset have an elevation in the Costimulation by the CD28 family pathway relative to proliferative (p = 0.0026) or normal-like (p=0.0001) patients (Figure 1c). In inflammatory patients, we see a correlation (R=-0.62, p=0.02) between DmRSS and baseline expression ofthe Costimulation by the CD28 family pathway (Figure 1d).
Conclusion:
Abatacept shows the most benefit for inflammatory patients. CD28-related pathways are elevated in inflammatory patients relative to fibroproliferative or normal-like patients. Importantly, we find that the extent of improvement in inflammatory patients is directly correlated to their baseline expression of the Costimulation by the CD28 familypathway. We show that baseline gene expression can predict differential response to biological intervention by directly modulating pathways pertinent to a drug’s mechanism of action. This data suggests that stratifying patients by baseline signatures may clarify the effects observed and act as a step towards precision medicine for future clinical practice.
To cite this abstract in AMA style:
Mehta B, Franks J, Yuan Y, Wang Y, Berrocal V, Wood T, Spino C, Fox D, Khanna D, Whitfield M. Machine-learning Classification Identifies a Subset of Patients That Improve on Abatacept via Modulation of a CD28-Related Pathway [abstract]. Arthritis Rheumatol. 2019; 71 (suppl 10). https://acrabstracts.org/abstract/machine-learning-classification-identifies-a-subset-of-patients-that-improve-on-abatacept-via-modulation-of-a-cd28-related-pathway/. Accessed .« Back to 2019 ACR/ARP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/machine-learning-classification-identifies-a-subset-of-patients-that-improve-on-abatacept-via-modulation-of-a-cd28-related-pathway/