Session Information
Session Type: Abstract Session
Session Time: 2:00PM-2:15PM
Background/Purpose: Skin inflammation and photosensitivity are common manifestations of cutaneous (CLE) and systemic lupus erythematosus (SLE), yet the mechanisms underlying heightened cell death and epidermal inflammation remain unclear. Non-lesional lupus keratinocytes (KCs) sustain an interferon (IFN)-rich, inflammatory profile even through several weeks of culture, suggesting possible epigenetic changes that drive this phenotype. Here we identify differential methylation of the Hippo signaling pathway as a novel regulator of apoptosis and photosensitive responses in lupus skin.
Methods: Genome-wide DNA methylation analysis was performed on extracted KC DNA from SLE patients and HC skin biopsies. Single-cell and bulk RNA sequencing was used to analyze RNA expression in cultured primary KCs from SLE patients and HC.In vivo protein and phospho-protein expression was assessed by immunofluorescent staining of HC and non-lesional lupus skin. A spontaneously immortalized KC line (N/TERTs) expressing a tetracycline inducible TEAD inhibitor (TEADi) that prevents colocalization of YAP-TEAD was used to evaluate the role of YAP signaling in apoptosis and UVB-mediated cell death. Apoptosis was analyzed via TUNEL staining and caspase 3/7 assay.
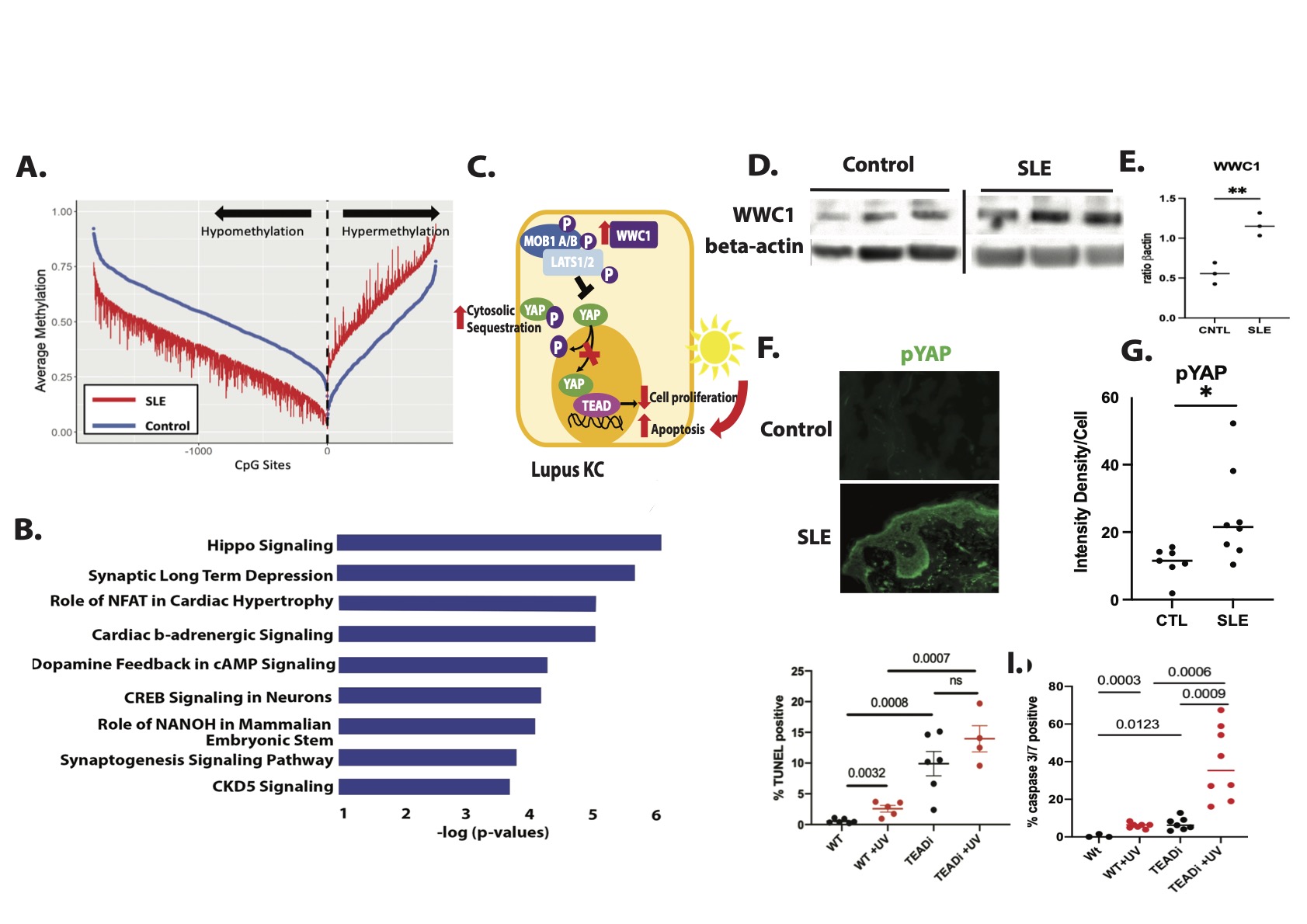
Results: Genome-wide DNA methylation analysis on cultured KC DNA from non-lesional, non-sun exposed skin biopsies of SLE patients and HC identified Hippo signaling as the top differentially-methylated pathway. We found significant hypomethylation of WWC1 in lupus KCs compared to control (Δβ=-0.17,P= 4.36 X 10-9) which promotes phosphorylation and thus cytosolic sequestration of YAP, attenuating TEAD-mediated transcription of pro-proliferative target genes. To determine functional relevance, we compared IFNγ or IFNα stimulated RNA-seq samples from the KCs used for our methylation studies. We found a negative correlation between IFN responses and methylation signatures. As confirmation, we analyzed Hippo regulated genes using single-cell RNA sequencing of nonlesional KC isolated from skin biopsies of lupus vs. HC and found downregulation of pro-proliferative YAP target genes such as SLP1, MYC, and AREG. Consistent with modulation of Hippo signaling as predicted by our methylation data, WWC1 expression was significantly increased in SLE vs. HC KC (p = 0.0142). Further, immunofluorescent microscopy of frozen non-lesional biopsies identified a significant increase in phosphorylated YAP in lupus skin compared to controls. TEADi induction in a spontaneously immortalized KC line resulted in a robust increase in UVB-mediated apoptosis as measured by TUNEL staining and caspase 3/7 activation, suggesting that the cytoplasmic retention of YAP observed in SLE KCs may contribute to enhanced UV-mediated KC apoptosis.
Conclusion: Collectively, our work describes a novel mechanistic paradigm in lupus KC in which hypomethylation of key regulators of the Hippo pathway restrict coactivation of TEAD transcriptional activity through cytoplasmic retention of YAP. This results in a propensity for apoptosis after UVB. Thus, modulation of the Hippo pathway could serve as a novel target for abrogation of photosensitivity in SLE and CLE patients.
To cite this abstract in AMA style:
Hile G, Coit P, Xu B, Estadt S, Martens J, Wasikowski R, Tsoi L, Iglesias-Bartolome R, Berthier C, Billi A, Gudjonsson J, Sawalha A, Kahlenberg J. Hippo Signaling Is a Novel Regulator of Apoptosis and Photosensitivity in Lupus Keratinocytes [abstract]. Arthritis Rheumatol. 2021; 73 (suppl 9). https://acrabstracts.org/abstract/hippo-signaling-is-a-novel-regulator-of-apoptosis-and-photosensitivity-in-lupus-keratinocytes/. Accessed .« Back to ACR Convergence 2021
ACR Meeting Abstracts - https://acrabstracts.org/abstract/hippo-signaling-is-a-novel-regulator-of-apoptosis-and-photosensitivity-in-lupus-keratinocytes/