Session Information
Date: Monday, November 9, 2020
Title: Pediatric Rheumatology – Clinical Poster III: SLE, Vasculitis, & JDM
Session Type: Poster Session D
Session Time: 9:00AM-11:00AM
Background/Purpose: Familial Hemophagocytic lymphohistiocytosis (fHLH) is an autosomal recessive, hyper-inflammatory, life-threatening disease. Macrophage activation syndrome (MAS) is also known as secondary HLH due to the clinical and biological similarities. MAS is increasingly recognized as a complication of systemic lupus erythematosus (SLE). This study compares the frequency of HLH genetic variants in childhood-onset SLE (cSLE) patients with and without MAS.
Methods: Our retrospective cohort was drawn from the Lupus Clinic at SickKids Hospital (Toronto) from 1987-2018. All participants met ACR, and/or SLICC SLE classification criteria. MAS diagnosis was based on expert physician diagnosis. The non-MAS cSLE comparator group was comprised of patients suspected monogenic SLE with whole genome sequencing (WGS). The non-MAS patients had young age of diagnosis (< 11y), consanguineous parents or multiple SLE affected first-degree relatives. Patients with MAS had paired-end whole exome sequencing (WES) by Illumina HiSeq 2500 platform (read depth 70-118X). Patients without MAS had paired-end WGS by Illumina HiSeq X platform (read depth 37-40X). Variant calling was completed with GATK and HAS and functionally annotated with ANNOVAR at The Centre for Applied Genomics (TCAG) in SickKids (annotation pipeline, v26.2, v.26.5). We restricted analyses to exonic regions in the following 16 HLH-genes (AP3B1, BLOC1S6, CD27, GATA2, ITK, LYST, MAGT1, NLRC4, PRF1, RAB27A, SH2D1A, SLC7A7, STX11, STXBP2, UNC13D, XIAP). Missense variants with MAF < 0.05 were prioritized. Ancestry was genetically inferred from multiethnic genotyping arrays (Illumina MEGA or GSA) using the ADMIXTURE software and the 1000 Genomes Project as reference. Otherwise, we used self-reported ethnicity according to Canada census categories. Demographic and baseline characteristics were extracted from the Lupus dataset, and analyzed using summary statistics. Medians and interquartile ranges (IQR) were calculated for non-normally distributed data and P-values were derived using two-tailed Fischer exact test.
Results: Our cohort consisted of 19 patients with MAS and 64 without MAS. The age of cSLE diagnosis was younger in the non-MAS cohort but the ethnicity distributions were no different between the two cohorts (Table). Eleven (58%) MAS and 39 (61%) non-MAS patients had exonic HLH variants; 6 (32%) MAS and 30 (47%) non-MAS had missense HLH variants. All these HLH variants were heterozygotes. There was no statistically significant difference in the number of HLH variants (exonic or missense) between MAS and non-MAS cohorts.
Conclusion: We did not observe a difference in the number of HLH exonic or missense variants in cSLE patients with MAS compared to those without MAS. Although this is the largest study of HLH variants in cSLE to date, our sample size had limited power to detect significant differences in rare variants which may have included variants associated with HLH. The non-MAS comparator cohort was likely enriched with rare pathogenic variants based on selection criteria for WGS. Future sequencing studies in larger cohorts of MAS and non-MAS cSLE patients are required to validate our findings.
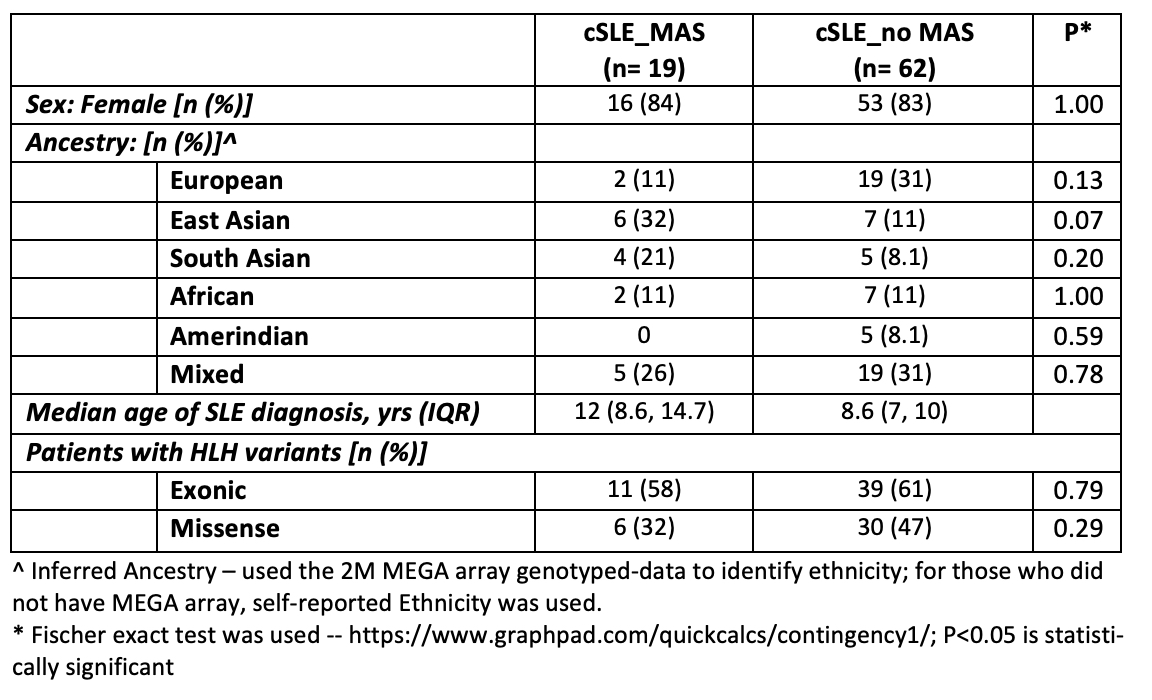 Table: Demographic and HLH variant comparison of cSLE patients with and without MAS
Table: Demographic and HLH variant comparison of cSLE patients with and without MAS
To cite this abstract in AMA style:
Lahiry P, Naumenko S, Liao F, Dominguez D, Knight A, Levy D, Misztal M, Ng L, Silverman E, Hiraki L. Hemophagocytic Lymphohistiocytosis (HLH) Gene Variants in Childhood-onset SLE (cSLE) with Macrophage Activation Syndrome (MAS) [abstract]. Arthritis Rheumatol. 2020; 72 (suppl 10). https://acrabstracts.org/abstract/hemophagocytic-lymphohistiocytosis-hlh-gene-variants-in-childhood-onset-sle-csle-with-macrophage-activation-syndrome-mas/. Accessed .« Back to ACR Convergence 2020
ACR Meeting Abstracts - https://acrabstracts.org/abstract/hemophagocytic-lymphohistiocytosis-hlh-gene-variants-in-childhood-onset-sle-csle-with-macrophage-activation-syndrome-mas/
