Session Information
Session Type: Abstract Submissions (ACR)
Background/Purpose: The aim of this study is to search for osteoarthritis (OA) biomarkers in human serum using an easy and fast approach and the validation of the potential biomarkers found with two independent methods.
Methods: The serum samples for the biomarker discovery experiment were obtained from 20 OA patients and 20 non-symptomatic controls. Samples were grouped into four pools of 10 samples each. The pools were subjected to a chemical sequential depletion protocol to reduce the high dynamic range of the proteins. Then, the proteomics comparison between OA and control sera was performed across two dimensional difference in-gel electrophoresis (2D-DIGE) experiment. The quantitative image analysis was performed using Same Spots software. For protein identification, gel spots were digested and analyzed by mass spectrometry (MALDI-TOF/TOF) and identified using Mascot with SwissProt knowledgebase.
Haptoglobin chains validation was performed with two different methods, immunoblotting and multiple reaction monitoring (MRM) technology with the QTRAP mass spectrometer.
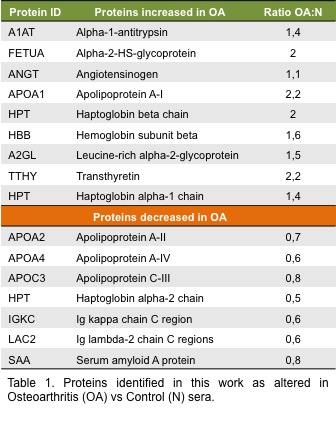
Results: We studied the combination of a chemical sequential depletion method combined with 2D-DIGE for the search of OA biomarkers in 40 serum samples. The analysis resulted in 46 spots significantly and reproducibly altered between OA and control samples (29 increased and 17 decreased). These 46 spots correspond to 14 different proteins, Table 1. The most interesting result was the modulation of the protein haptoglobin (HPT), three different spots were opposite modulated and were identify as the three chains of haptoglobin. In human, HPT exists in two allelic forms that produce three known phenotypes. The presence of the different alpha chains in patients depends on which phenotype their express. Interestingly, HPT beta and HPT alpha-1 chains were increased in OA sera whereas HPT alpha-2 chain was decreased in OA sera versus control. Using western blot analysis and MRM mass spectrometry technique on 30 new individual samples from OA and control donors (15 from each condition), we confirmed the different modulations of the alpha and beta HPT chains.
Conclusion: We were able to identified 16 protein forms altered in the disease (9 increased and 7 decreased), and we verified for the first time the OA- dependent alteration of the haptoglobin chains. The haptoglobin protein provides two different OA biomarkers easily to measure in serum samples at the same time.
Disclosure:
C. Fernandez-Costa,
None;
V. Calamia,
None;
P. Fernandez-Puente,
None;
J. Mateos,
None;
B. Rocha,
None;
L. Lourido,
None;
J. L. Capelo,
None;
C. Ruiz-Romero,
None;
F. J. Blanco,
None.
« Back to 2013 ACR/ARHP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/haptoglobin-chains-as-potential-biomarkers-in-serum-of-osteoarthritis-disease/