Session Information
Session Type: Poster Session (Monday)
Session Time: 9:00AM-11:00AM
Background/Purpose: Fragmented cell-free DNA (cfDNA) is released into blood circulation as results of damage or death of peripheral blood cells as well as organ tissues, and it is significantly increased in those of cancer patients to be used in monitoring disease activities. CfDNA was also reported as a potential biomarker in autoimmune diseases, such as rheumatoid arthritis and systemic lupus erythematosus.
The cfDNA sequence is identical to that of genomic DNA, since the DNA of each cell type possesses specific methylation patterns. In eosinophils, methylation levels of the loci, interleukin 5 receptor alpha unit (IL5RA) and interleukin 4 (IL4), were reported to be lower as compared to other blood cells.
The purpose of this study is to evaluate the correlation between serum cfDNA and clinical disease activities in patients with Eosinophilic granulomatosis with polyangiitis (EGPA), and to examine tissue specific methylation patterns of cfDNA.
Methods: The study group included 24 patients with ANCA associated vasculitis (AAV), 8 patients with EGPA, 7 with microscopic polyangiitis (MPA) and 9 with granulomatosis with polyangiitis (GPA). CfDNA was extracted from serum using Qiamp MinElute cfDNA kit (QIAGEN). By quantitative real-time PCR, the concentration of cfDNA was measured twice retrospectively; before immunosuppressive therapy and remission period. CfDNA from 10 healthy controls (HC) was also measured. Then, we evaluated the correlation between cfDNA and the clinical disease activity, using Birmingham vasculitis score (BVAS) and C reactive protein (CRP).
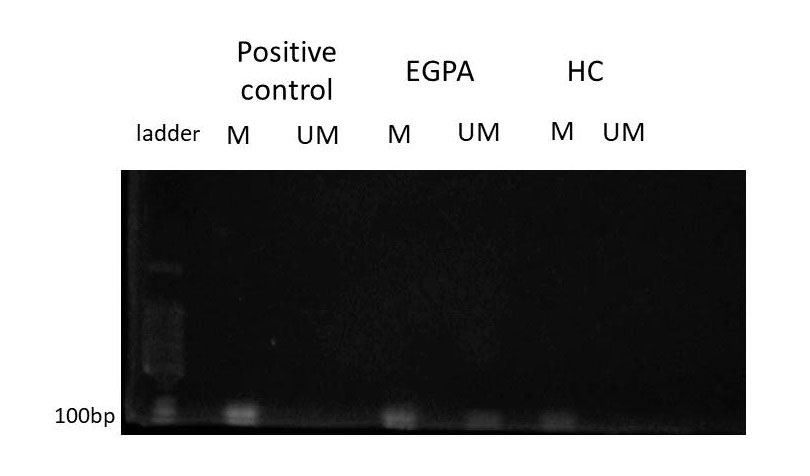
Next, cfDNA from patients with EGPA and HC was treated with bisulfite to convert unmethylated cytosines to uracils. Methylation specific PCR was performed to evaluate two loci, IL5RA and IL4.
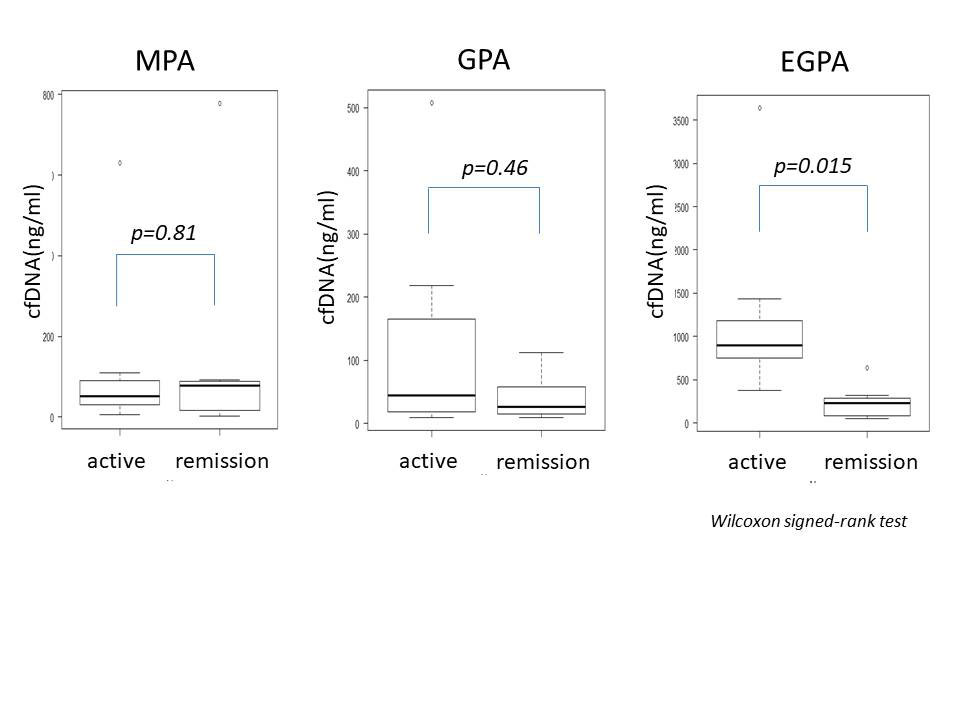
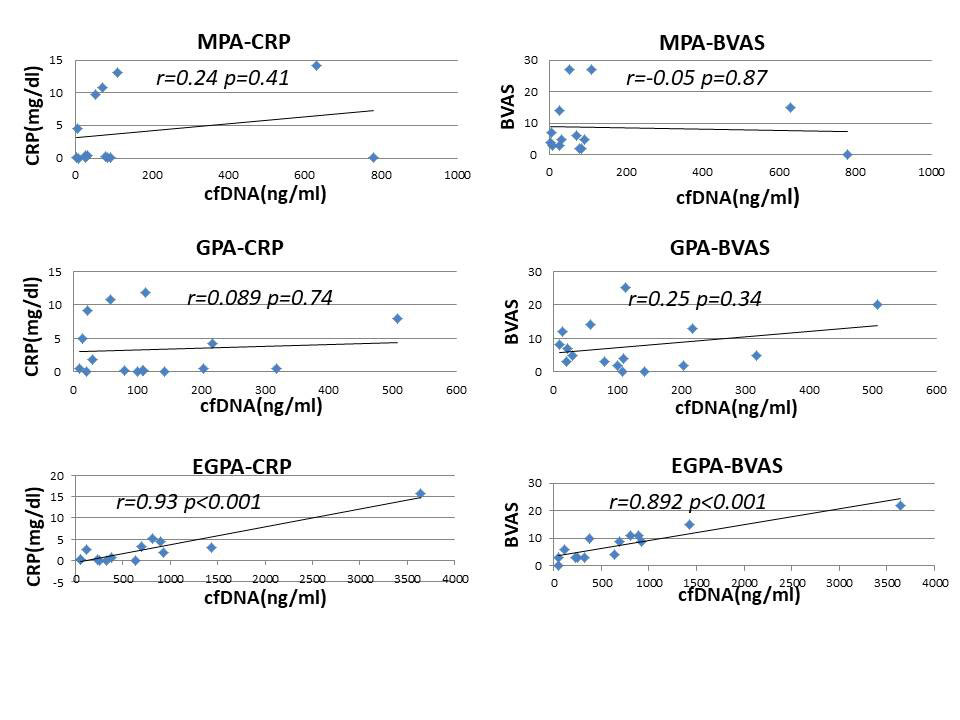
Results: The concentration of cfDNA in AAV was higher than HC (p< 0.001) before treatment, and cfDNA in EGPA was significantly higher than the other AAV (p< 0.001). After introduced immunosuppressive therapy, cfDNA was significantly decreased in EGPA (p=0.015), but not changed in MPA (p=0.81) and GPA (p=0.46) (fig.1). In patients with EGPA, cfDNA was strongly correlated with BVAS (EGPA; r=0.89, p< 0.001, MPA; r=-005, p=0.87 and GPA; r=0.25, p=0.34, respectively). CfDNA also strongly correlated with CRP in EGPA (EGPA; r=0.92, p< 0.001, MPA; r=0.24, p=0.4 and GPA; r=0.09, p=0.74, respectively) (fig.2). Methylated DNA of IL5RA and IL4 was detected in both EGPA and HC, while those of un-methylated DNA was detected in EPGA (fig.3).
Conclusion: CfDNA is expected to be a useful biomarker for EGPA, representing the lower methylation of IL5RA and IL4 loci in eosinophils from patients with EPGA .
positive control: Methylated HeLa gDNA
To cite this abstract in AMA style:
Hashimoto T, Yokoyama Y, Furukawa T, Yoshikawa T, Azuma N, Yoshida K, Hashiramoto A, Matsui K. Circulating Cell Free DNA Released from Eosinophils Is a Practical Biomarker in Patients with Eosinophilic Granulomatosis with Polyangiitis [abstract]. Arthritis Rheumatol. 2019; 71 (suppl 10). https://acrabstracts.org/abstract/circulating-cell-free-dna-released-from-eosinophils-is-a-practical-biomarker-in-patients-with-eosinophilic-granulomatosis-with-polyangiitis/. Accessed .« Back to 2019 ACR/ARP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/circulating-cell-free-dna-released-from-eosinophils-is-a-practical-biomarker-in-patients-with-eosinophilic-granulomatosis-with-polyangiitis/