Session Information
Date: Monday, November 13, 2023
Title: Abstracts: Systemic Sclerosis & Related Disorders – Basic Science
Session Type: Abstract Session
Session Time: 4:00PM-5:30PM
Background/Purpose: Interstitial lung disease (ILD) is present in up to 90% of patients with systemic sclerosis (SSc) and the leading cause of SSc-related mortality. SSc-ILD is categorized radiographically and pathologically in two subtypes: non-specific interstitial pneumonia (NSIP) and usual interstitial pneumonia (UIP). Epidemiologic studies show SSc-UIP has a significantly worse prognosis than SSc-NISP and does not respond to immunomodulating therapy. However, despite these clinical differences, little is known regarding the molecular profiles of the two SSc-ILD subtypes, making biomarker discovery and treatment development challenging. The objective of this study is to define the cellular and molecular profiles underpinning SSc-NSIP and SSc-UIP compared to idiopathic pulmonary fibrosis (IPF).
Methods: Single nuclei RNA sequencing (snRNAseq) was performed on frozen explanted lung tissue of 20 age and sex matched individuals including 10 SSc (5 NSIP and 5 UIP), 5 IPF, and 5 healthy controls (HC). Nuclei were isolated, multiplexed, and sequenced using the 10x genomics 3′ v3 platform. Data were processed using standard CellRanger and Seurat pipelines and SoupX, Harmony, and Demuxlet. Cell proportions between conditions were compared using Kruskall Wallis test and differential expression was performed using DESeq2. Unsupervised hierarchical clustering of DEGs was used to examine similarities and differences between SSc-ILD subtypes within cell subsets.
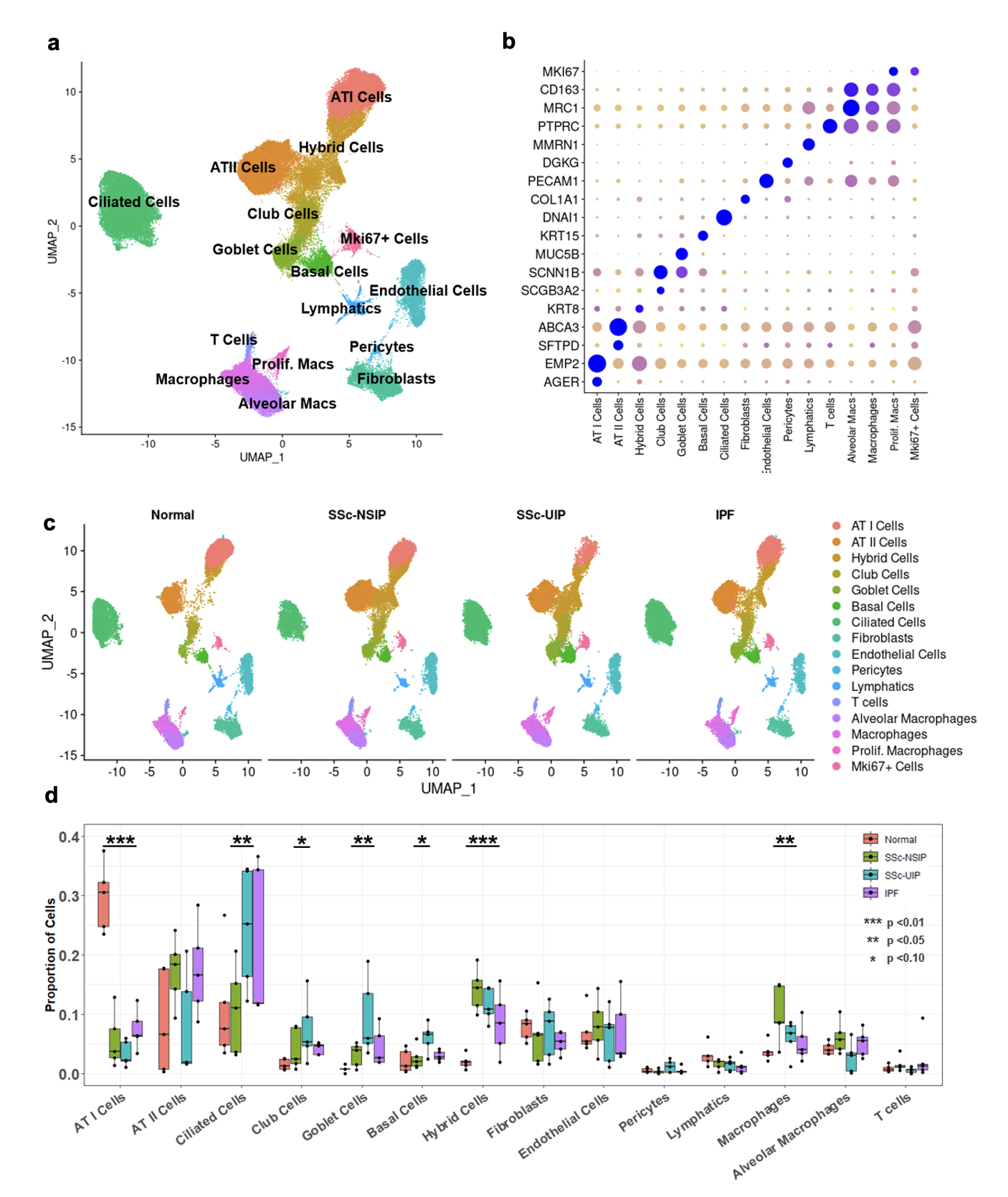
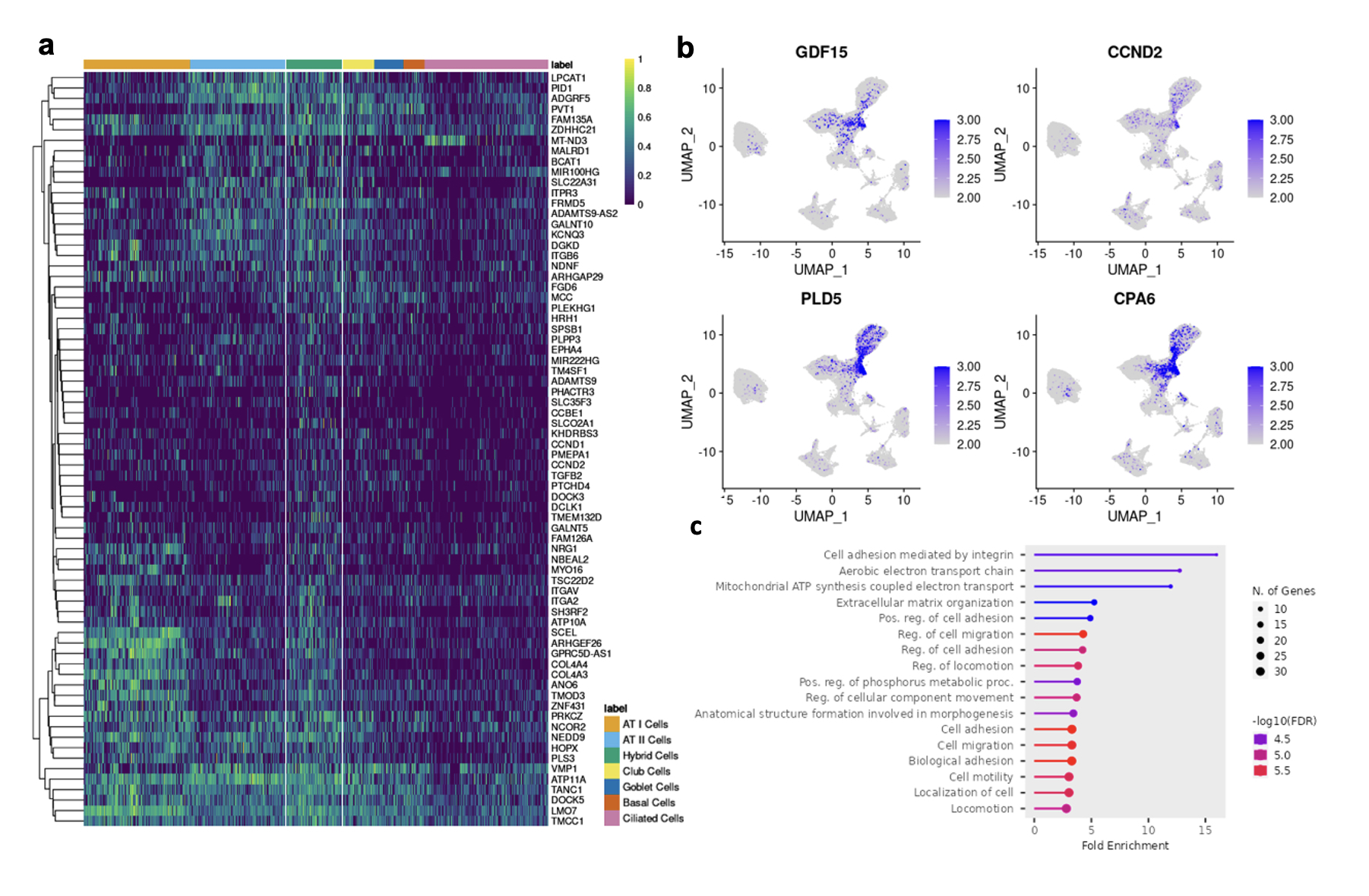
Results: Following QC filtering, a total of 92,128 nuclei, including 20,738 SSc-NSIP, 24,012 SSc-UIP, 21,122 IPF, and 26,256 healthy nuclei, were used in downstream analysis. Clustering revealed 16 distinct cell types (Fig 1a) labeled using canonical gene markers (Fib 1b). Cell composition analysis revealed a profound loss of ATI cells in the diseased lungs compared to HC (p< 0.01) and an enrichment of other epithelial cells including ciliated and goblet cells in SSc-UIP and IPF. Macrophages were uniquely enriched in SSc-NSIP (Fig 1c-d). A “Hybrid” epithelial cell was identified that was unique to diseased lung and significantly increased in both SSc-ILD subtypes. DEG analysis of this cell group revealed 133 genes of which some were shared with other epithelial cells, such as ATI and ATII, while others were unique to the population (Fig 2a). Notable genes included markers of cellular senescence, i.e., GDF15 and CCND2, and previously described markers of “aberrant cells” in IPF, i.e., PLD5 and CPA6 (Fib 2b). Pathway analysis revealed upregulation of extracellular matrix organization and cell adhesion and migration processes. Comparing SSc-ILD subtypes, epithelial cells demonstrated a distinct expression pattern between SSc-NSIP and SSc-UIP which was redemonstrated in ATI cells, Club cells, and Basal Cells, while other epithelial subsets showed a more similar pattern of expression between the SSc-ILD subtypes.
Conclusion: This study provides novel insight to the molecular makeup of the SSc-ILD subtypes and demonstrates that SSc-NSIP and SSc-UIP have differing cell compositions but have both shared and distinct molecular pathways that also overlap with IPF. Ongoing analyses will further elucidate the molecular profiles underpinning the SSc-ILD subtypes.
To cite this abstract in AMA style:
Yang M, Deiter F, Flynn E, Neely J, Lee S, Greenland J, Sirota M, Wolters P. Cell Specific Molecular Profiling of Scleroderma Associated Interstitial Lung Disease Subtypes [abstract]. Arthritis Rheumatol. 2023; 75 (suppl 9). https://acrabstracts.org/abstract/cell-specific-molecular-profiling-of-scleroderma-associated-interstitial-lung-disease-subtypes/. Accessed .« Back to ACR Convergence 2023
ACR Meeting Abstracts - https://acrabstracts.org/abstract/cell-specific-molecular-profiling-of-scleroderma-associated-interstitial-lung-disease-subtypes/