Session Information
Session Type: Poster Session A
Session Time: 9:00AM-11:00AM
Background/Purpose: Glucocorticoids (GC), utilised in SLE for their broad immunosuppressive actions, predominantly mediate these effects by interaction with the cytoplasmic GC receptor (GR) to modulate gene transcription. Interactions of GC with many pathways in SLE, including the type 1 interferon (IFN) program, are poorly understood, with some evidence that IFN program is GC resistant. Minimising GC exposure is a treatment goal in SLE, however evidence-based treatment guidelines are lacking and no pharmacodynamic measure has been established. We aimed to evaluate GC regulated genes for their potential to be used as biomarkers of GC exposure in SLE.
Methods: A previously described set of 80 GC induced genes (Hu et al, Arthritis and Rheumatology, 2018) was measured using Fluidigm Biomark HD in peripheral blood mononuclear cells (PBMC) from patients meeting ACR criteria for SLE. Public data sets GSE49454 and GSE 88884 were accessed; clinical data correlating to GSE88884 were provided by Eli Lily. IFN status was determined using a validated 4 gene signature (IFI44, IFI44L, IFI27, RSAD2). RNAseq was performed on RNA extracted from healthy donor PBMC (n=4) treated with dexamethasone 10-7M (DEX) and/or IFN 1000IU/mL. The HALLMARK_INTERFERON_ALPHA_RESPONSE gene set was extracted from the Broad Institute Molecular Signature Database.
Results: 80 GC-regulated genes were analysed in our cohort (n=18) and GSE49454 (n=62). Five genes (VSIG4, ALOX15B, CD163, AMPH, IL1R2) correlated with daily oral GC dose in both cohorts; these were combined to make a 5-gene GC signature (5GGCS) and this was analysed in GSE88884 (n=1,756). Although a dose-dependent association of 5GGCS expression with GC doses was observed (p < 0.0001) (Fig 1A), high interpatient variation (Fig 1B) meant GC signature had only modest ability to distinguish patients taking/not taking GC overall (AUC = 0.69) (Fig 1C). We next examined whether IFN status impacted on GC signatures in SLE. In GSE88884 there was no effect of GC dose on ISG expression. In contrast, we observed a stronger correlation of 5GGCS with GC dose in IFN low patients (R=0.42, p< 0.0001) compared to IFN high patients (R=0.29, p< 0.0001) and IFN low patients had increased 5GGCS expression compared to IFN high patients matched for GC dose (p < 0.05), suggesting GC-induced genes are suppressed by IFN. RNAseq on IFN and DEX-treated PBMC confirmed that IFN significantly altered the expression of 29/80 GC-regulated genes, while DEX had minimal impact on ISG expression. We identified 61 genes regulated by DEX but not affected by IFN. Only 9/61 correlated with GC dose in GSE88884. Combined signatures using these genes showed modest ability to distinguish patients taking/not taking GC (AUC=0.63) (Figure 2), with slightly better performance for negatively regulated genes when prednisolone ≥ 25 mg/day (AUC=0.71).
Conclusion: A signature for GC exposure was identified, but interpatient variation and effects of IFN status may limit application in IFN-driven diseases such as SLE. These data confirm insensitivity of IFN-regulated genes to GC, but substantial impact of IFN on GC induced genes, strongly supporting the concept that IFN plays a role in GC resistance in SLE.
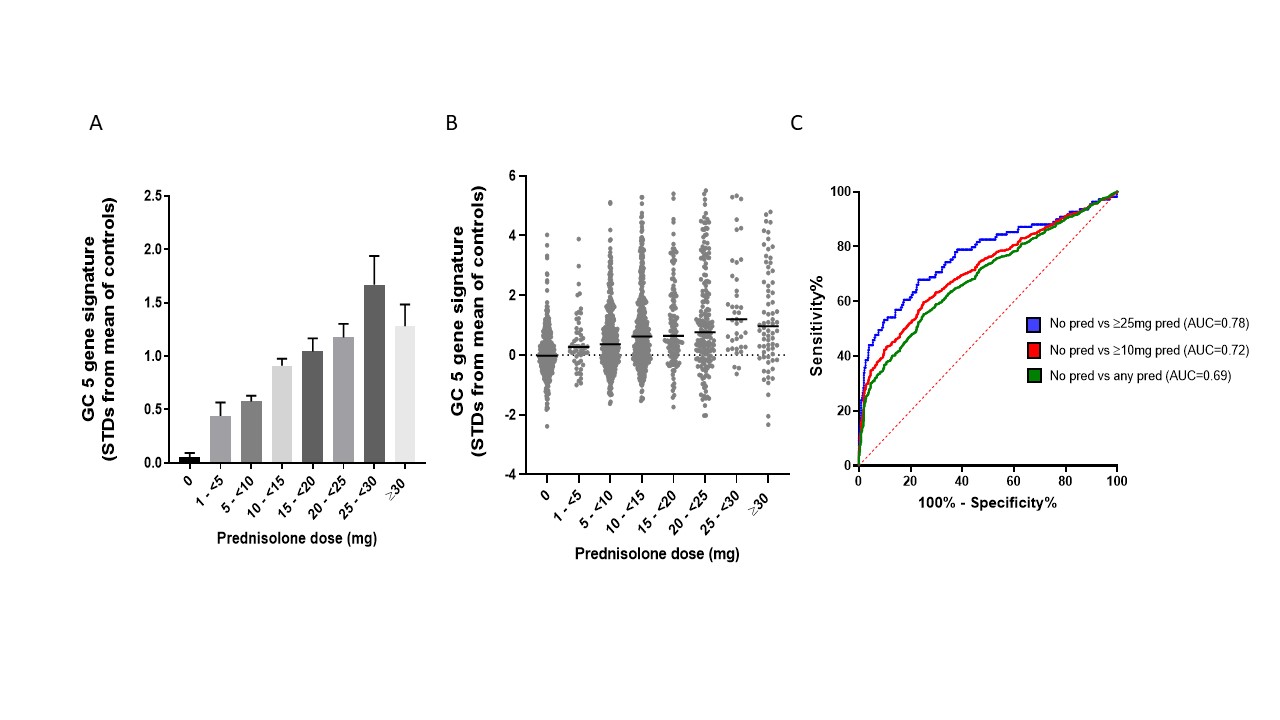 Figure 1. Correlations between 5-gene GC signature and prednisolone (pred) dose in dataset GSE888884. Mean GC signature scores increased with increasing GC dose (A, p for trend < 0.0001) but exhibited large interpatient variation (B) and only modest benefit in differentiating patients taking/not taking prednisolone (C).
Figure 1. Correlations between 5-gene GC signature and prednisolone (pred) dose in dataset GSE888884. Mean GC signature scores increased with increasing GC dose (A, p for trend < 0.0001) but exhibited large interpatient variation (B) and only modest benefit in differentiating patients taking/not taking prednisolone (C).
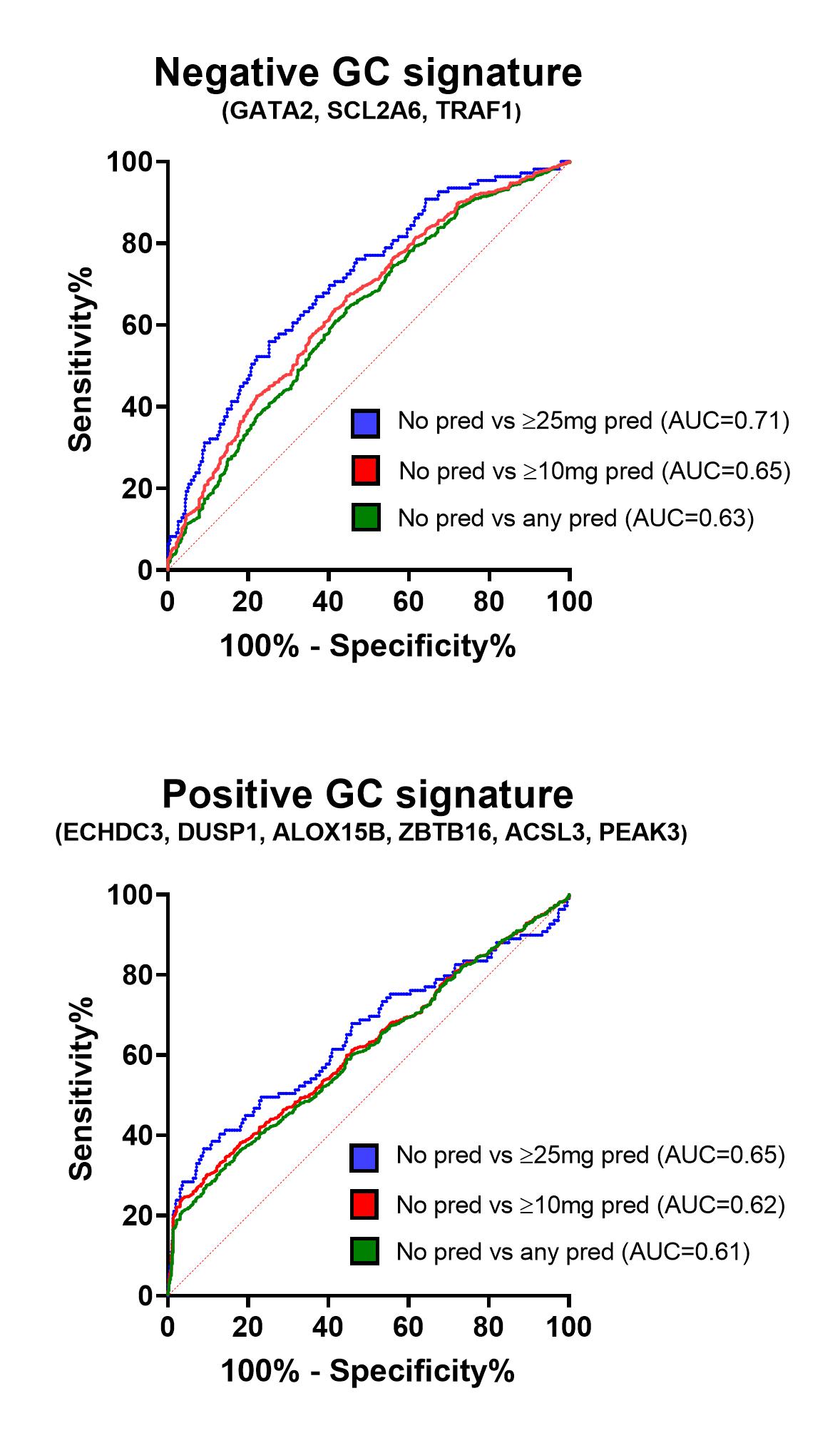 Figure 2. ROC curve analysis for negative and positive GC gene signatures to distinguish patients taking/not taking GC prednisolone (pred).
Figure 2. ROC curve analysis for negative and positive GC gene signatures to distinguish patients taking/not taking GC prednisolone (pred).
To cite this abstract in AMA style:
Northcott M, Gearing L, Nim H, Nataraja C, Jones S, Morand E. Towards a Glucocorticoid Exposure Signature in SLE: Effects of Type I Interferon [abstract]. Arthritis Rheumatol. 2020; 72 (suppl 10). https://acrabstracts.org/abstract/towards-a-glucocorticoid-exposure-signature-in-sle-effects-of-type-i-interferon/. Accessed .« Back to ACR Convergence 2020
ACR Meeting Abstracts - https://acrabstracts.org/abstract/towards-a-glucocorticoid-exposure-signature-in-sle-effects-of-type-i-interferon/
