Session Information
Date: Tuesday, November 14, 2023
Title: (2257–2325) SLE – Diagnosis, Manifestations, & Outcomes Poster III
Session Type: Poster Session C
Session Time: 9:00AM-11:00AM
Background/Purpose: Epidemiologists, health services researchers, and health outcome investigators have begun utilizing real-world data (RWD) to gain valuable insights into disease patterns, treatment outcomes, and healthcare utilization and economics. Similarly, biopharmaceutical and medical device manufacturers utilize RWD to identify medical gaps, enhance patient care, and drive innovative therapeutic interventions. However, a significant portion (~80%) of essential clinical data is trapped in unstructured text within clinical notes, thereby remaining inaccessible in existing RWD for systemic lupus erythematosus (SLE).
Methods: To address this challenge, we leveraged the cutting-edge Natural Language Processing (NLP) model, ChatGPT 4.0, to analyze clinical notes to identify potential diagnosis of SLE. We evaluated:1) the adequacy of unstructured clinical notes in capturing essential clinical details (i.e., 22 clinical and immunologic variables) to support meeting the 2019 European League Against Rheumatism and American College of Rheumatology (EULAR/ACR) classification criteria for SLE; 2) the efficacy of the ChatGPT model in identifying those variables outlined in the EULAR/ACR criteria.
De-identified medical records were obtained from Temple University Health System’s EHR system (Epic), comprised patients aged 18 years or above, with at least at least one rheumatology visit and one ICD-10-CM code of M32.* (SLE) between 1/1/2012 and 5/31/2022. Using a Python script, we conducted an automatic search for various mentions of SLE in rheumatology visit notes. We then identified those with a positive anti-nuclear antibody (ANA) test. A thorough manual review of the rheumatology notes were conducted to determine if patients met the EULAR/ACR classification criteria. Subsequently, we employed ChatGPT 4.0 to extract the EULAR/ACR clinical variables.
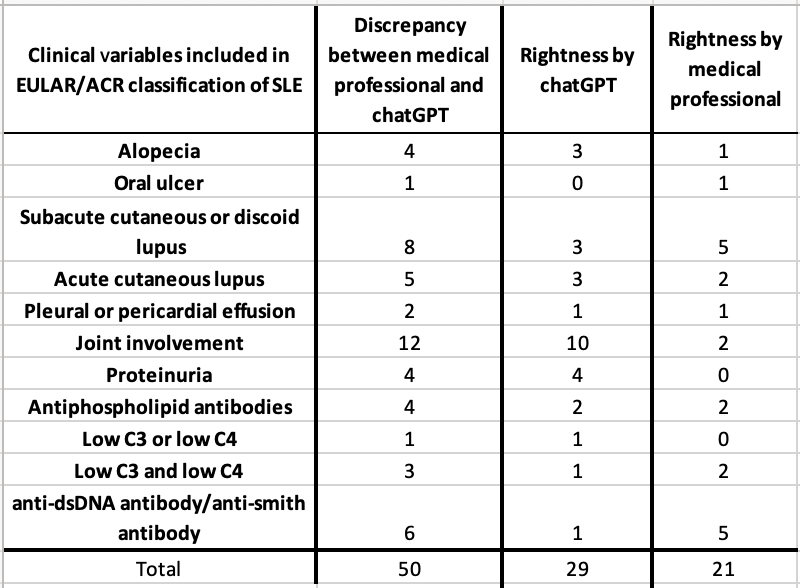
Results: The initial Python search yielded 16,124 rheumatology notes from 445 unique patients. The follow-up search for positive ANA tests returned 1,147 rheumatology notes from 187 unique patients. The manual review by trained medical professionals revealed that 116 of the 187 patients met the EULAR/ACR criteria. There was a remarkably high level of agreement between the assessments made by ChatGPT 4.0 and those made by trained medical professionals (830 out of 880 clinical and immunologic variables across 40 randomly selected patients). Among the 50 annotations of disagreement (see Table 1), ChatGPT’s interpretations were found to be accurate in 29 cases. In some cases, the differentiation between acute cutaneous lupus vs. subacute cutaneous lupus was not clearly documented in the visit notes. In other cases, ChatGPT and human annotators had to infer the classification based on certain patient-reported symptoms like rash and photosensitivity.
Conclusion: Our findings confirm that unstructured clinical notes contain sufficient clinical details into SLE patient care, including 22 clinical and immunologic variables specified in the established classification criterion for SLE. Moreover, the utilization of state-of-the-art NLP techniques has significant promise in identifying those with SLE, potentially enhancing existing SLE research using RWD.
To cite this abstract in AMA style:
Yao L, Gebru L, Aweke B, Patel J, Vina E, Fleece D, Wu H. Leveraging ChatGPT for Real-World Systematic Lupus Erythematosus Data Curation from Electronic Health Records: A Feasibility Study [abstract]. Arthritis Rheumatol. 2023; 75 (suppl 9). https://acrabstracts.org/abstract/leveraging-chatgpt-for-real-world-systematic-lupus-erythematosus-data-curation-from-electronic-health-records-a-feasibility-study/. Accessed .« Back to ACR Convergence 2023
ACR Meeting Abstracts - https://acrabstracts.org/abstract/leveraging-chatgpt-for-real-world-systematic-lupus-erythematosus-data-curation-from-electronic-health-records-a-feasibility-study/