Session Information
Date: Monday, November 8, 2021
Title: Spondyloarthritis Including PsA – Treatment Poster II: Psoriatic Arthritis I (1329–1363)
Session Type: Poster Session C
Session Time: 8:30AM-10:30AM
Background/Purpose: Guselkumab (GUS), an anti-IL-23 p19-subunit monoclonal antibody, demonstrated efficacy vs placebo (PBO) in reducing signs and symptoms of PsA in the phase-3 DISCOVER-1 & -2 studies.1,2 This study evaluated gene expression in the blood of PsA patients (pts) in DISCOVER-1 & -2 and the impact of GUS on the expression of these genes.
Methods: Pts were treated with GUS 100 mg every 4 weeks (Q4W); GUS 100 mg at Week (W) 0, W4, then Q8W; or matching PBO. Whole transcriptome profiling by RNA-sequencing was performed using the Novaseq platform on blood samples obtained from a subset of 673 pts with PsA at baseline across the 2 DISCOVER studies, as well as from 21 demographically (age, sex, ethnicity) matched healthy controls procured independently of the clinical program. A subgroup (N=227), selected based on having baseline characteristics (demographics, disease activity, medication use) representative of the overall cross-study PsA population, also had serial blood samples (W0/W4/W24) evaluated. Significance of differentially expressed genes (DEGs) between PsA and healthy controls was defined by a false discovery rate < 0.05 based on a log-linear model using edgeR. Top genes were defined by significance and |logFC| >1. For cell type analysis, genes that changed with GUS treatment were tested for enrichment using Cibersort. Gene enrichment scores were calculated using Gene Set Variation Analysis.
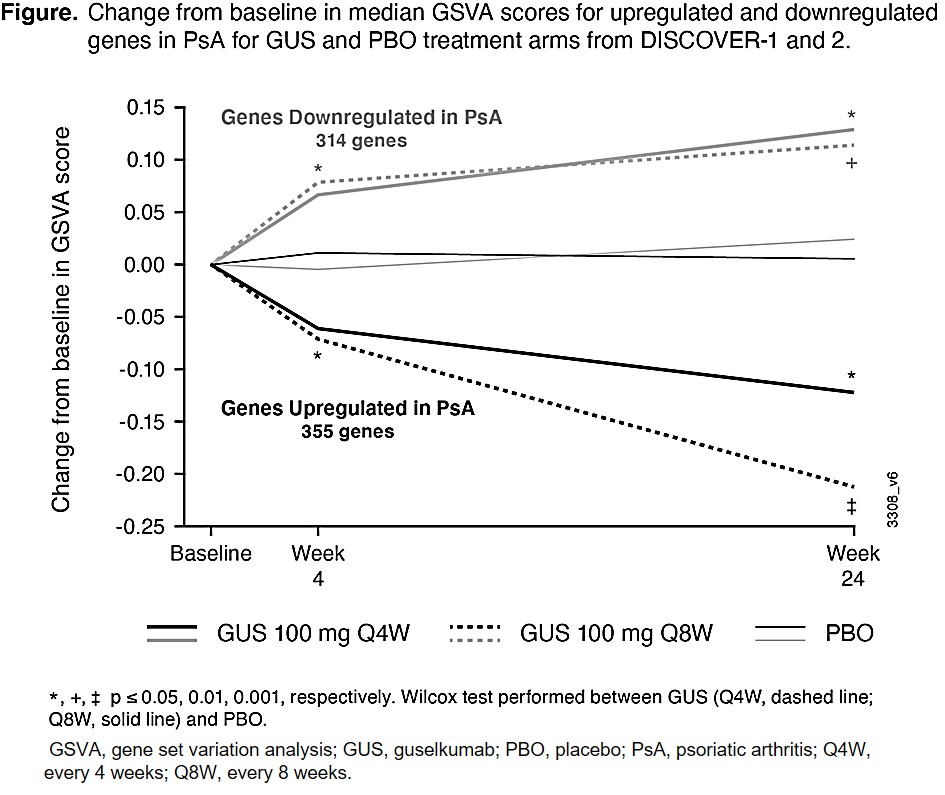
Results: To define disease genes, we compared genes at baseline in pts with active PsA vs healthy control whole blood transcriptomes and detected 355 upregulated and 314 downregulated (top genes shown in Table), defined here as core disease genes. Upregulated genes were largely related to neutrophils, monocytes, macrophages, and extracellular matrix, whereas downregulated genes were related to T cells. The upregulated disease genes were significantly decreased and the downregulated disease genes were significantly increased by GUS treatment vs PBO at W4 and W24 (Figure). Upon stratification by ACR 20 response, changes in core disease gene expression from W0 were statistically significant among responders, but not in nonresponders, at W4 and W24 (data not shown). We then performed the second differential expression analysis comparing baseline to W4 and W24 for both PBO and GUS treatment arms to define genes that changed with treatment over time. At W4 and W24, we found many DEGs from baseline with GUS treatment and none with PBO. These included genes related to B-, T-, NK-, and plasma cells (increased by GUS) and neutrophils, monocytes, eosinophils, and macrophages (decreased by GUS), suggestive of a partial normalization of immune cell composition in whole blood.
Conclusion: Using whole transcriptome profiling, we detected DEGs in blood samples obtained from PsA pts vs healthy controls, suggesting a dysregulation of immune cell profiles in PsA. The majority of these disease-associated genes were modulated by GUS, with directionality toward a normalization of whole blood transcriptomic signatures.
1. Deodhar A et al. Lancet. 2020;395:1115
2. Mease P et al. Lancet. 2020;395:1126
To cite this abstract in AMA style:
Siebert S, Sweet K, Ritchlin C, Hsia E, Kollmeier A, Xu X, Song Q, Miron M. Guselkumab Treatment Modulates Core Psoriatic Arthritis Gene Expression in Two Phase 3 Clinical Trials (DISCOVER-1 and -2) [abstract]. Arthritis Rheumatol. 2021; 73 (suppl 9). https://acrabstracts.org/abstract/guselkumab-treatment-modulates-core-psoriatic-arthritis-gene-expression-in-two-phase-3-clinical-trials-discover-1-and-2/. Accessed .« Back to ACR Convergence 2021
ACR Meeting Abstracts - https://acrabstracts.org/abstract/guselkumab-treatment-modulates-core-psoriatic-arthritis-gene-expression-in-two-phase-3-clinical-trials-discover-1-and-2/