Session Information
Session Type: Poster Session (Tuesday)
Session Time: 9:00AM-11:00AM
Background/Purpose: Circulating cell-free mitochondrial DNA (mtDNA) is released during non-apoptotic processes such as NETosis. Increased levels of mtDNA are associated with autophagic dysfunction and inflammasome activation. In critically ill patients mtDNA levels correlate with mortality. In Granulomatosis with Polyangiitis (GPA) there is evidence that innate immune system activation is involved in pathogenesis and propagation of disease. The role of autophagic dysfunction and subsequent effects on the inflammasome as it relates to mtDNA has yet to be robustly investigated in GPA. The aim of this study was determine whether increased circulating mtDNA levels correlate with disease activity in GPA.
Methods: Plasma samples and clinical data from the Wegener’s Granulomatosis Etanercept Trial were utilized. Cell-free mtDNA levels were measured in banked plasma by applying a SYBR Green-qPCR assay using a PRISM 7300 sequence detection system (Applied Biosciences). All mtDNA level data were log-transformed. Samples were categorized based on disease activity and severity. The primary aim was to determine if mtDNA levels correlated with disease activity as defined by BVAS/WG. Samples were classified as pre-flare, flare, or remission, with flare further subcategorized into severe or non-severe
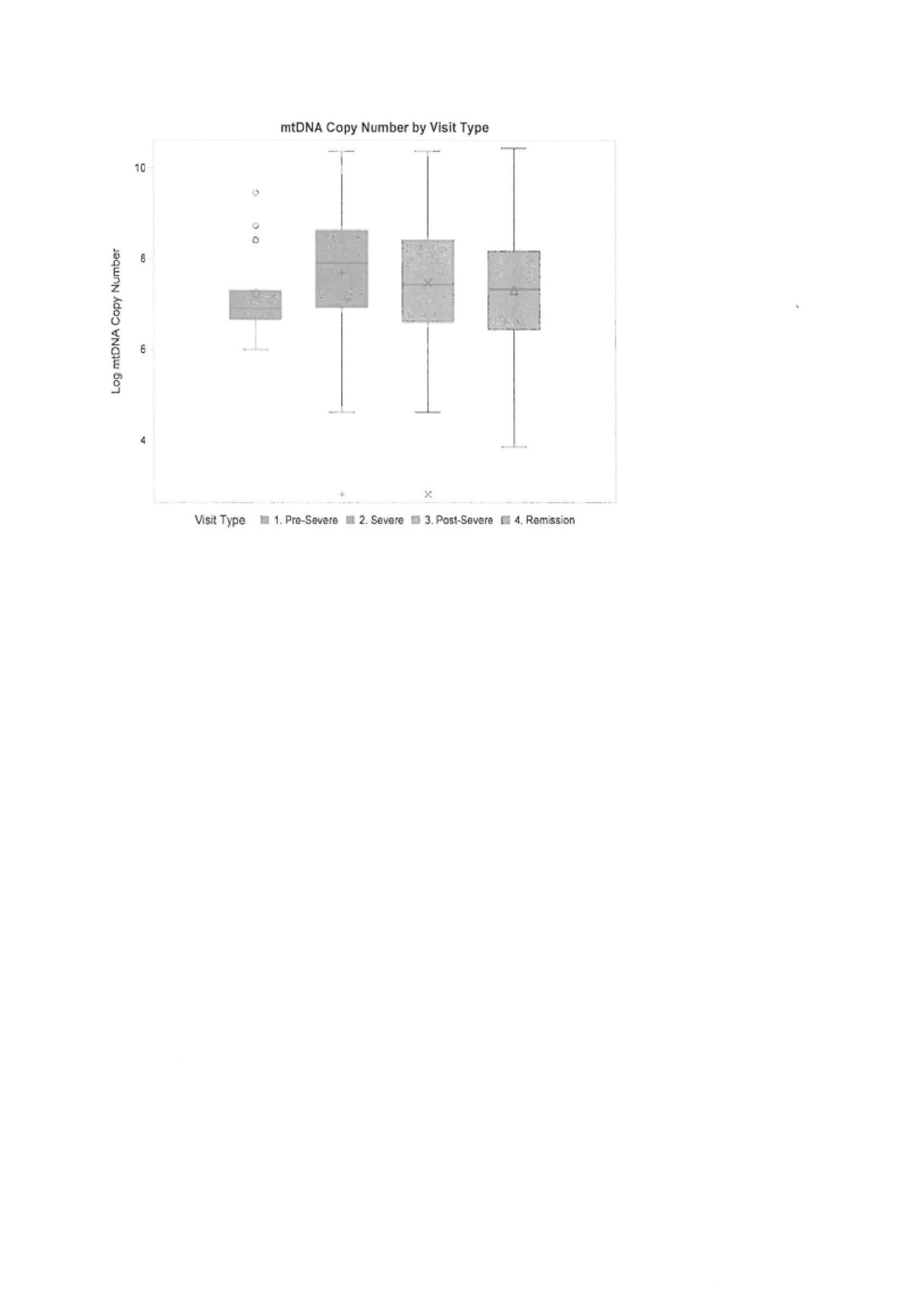
Results: 100 of the 180 participants in WGET had available plasma at 2 or more visits, and were data from these patients and sample were used for this analysis. No significant differences in patient demographics were observed between those with and without plasma. There was no difference in mtDNA copy number across visit types (pre-flare, flare, and remission). While mtDNA level was not a significant predictor of disease activity in univariate models, there was trend within severe flare: an increase in mtDNA level was indicative of increased odds of severe flare (Odds Ratio = 1.32; (0.78 – 2.22) In paired analysis (n=40), mean mtDNA level was significantly higher during a severe flare as compared to remission (M=7.75 ± 1.47, M=7.36 ± 1.17, respectively; p=.038). In a subset of the paired analysis (n=13), restricted to pre- and post-severe flare observations, mean mtDNA level was significantly higher 3 months following the flare as compared to pre-flare in relation to an index severe flare (7.83 ± 1.00 vs 7.25 ± 1.07, p=.052). Mean mtDNA between pre-severe flare and severe flare (n=9) was not significantly different (7.23 ± 1.15 vs 7.76 ± 1.32, p=.185).
Conclusion: This exploratory study suggests that mtDNA levels may be elevated in states of high disease activity among patients with GPA. However, firm conclusions are limited by small sample size. These findings suggest a role for deregulated autophagy in GPA. Finally, mtDNA level, a non-invasive and rapidly performed test, may potentially serve as a useful biomarker for disease activity in GPA, although additional examination in a larger cohort of patients is warranted to determine its utility and sensitivity.
To cite this abstract in AMA style:
Lally L, Sattui S, Finik J, Choi A, Rhee R, Merkel P, McCune W, St. Clair E, Specks U, Stone J, Spiera R. Cell-free Mitochondrial DNA Levels in Granulomatosis with Polyangiitis [abstract]. Arthritis Rheumatol. 2019; 71 (suppl 10). https://acrabstracts.org/abstract/cell-free-mitochondrial-dna-levels-in-granulomatosis-with-polyangiitis/. Accessed .« Back to 2019 ACR/ARP Annual Meeting
ACR Meeting Abstracts - https://acrabstracts.org/abstract/cell-free-mitochondrial-dna-levels-in-granulomatosis-with-polyangiitis/