Session Information
Session Time: 6:00PM-7:00PM
Background/Purpose: PFAPA (Periodic Fever, Aphthous Stomatitis, Pharyngitis, and cervical Adenitis) is the most common recurrent fever syndrome, yet a specific diagnostic biomarker is lacking. This absence leads to diagnostic delays, unnecessary invasive testing, and impaired quality of life. Our objective was to identify molecular mechanisms of PFAPA flares by identifying differentially expressed genes (DEGs) and proteins that distinguish flare from non-flare states, focusing on the salivary matrix as a potentially non-invasive sample source.
Methods: Paired blood and saliva samples were collected from 10 PFAPA patients fulfilling stringent inclusion criteria during both flare and non-flare states. Bulk RNA sequencing was performed on blood. Differentially-expressed genes were identified using standard pipelines (quality control, alignment to hg39 via Spliced Transcripts Alignment to a Reference-STAR, quantification, and DESeq2 analysis), followed by pathway enrichment using gene set enrichment analysis in python (GSEApy). Salivary proteomics data were analyzed using the Wilcoxon Signed-Rank Test. Semantic-similarity analysis was used to compare enriched Gene Ontology (GO) categories between the salivary proteome and blood transcriptome. Statistical significance was defined as p < 0.05 after FDR adjustment.
Results:
The median age of enrollment during flare was 100 (IQR: 78–134) months, with a maximum temperature of 104.9 (IQR 103.4 — 105.2) degrees Fahrenheit. RNA-Seq Gene Set Enrichment Analysis (GSEA) in the blood identified enrichment in key inflammatory pathways, including IFN signaling (Type I IFN, IFN-gamma), complement activation, neutrophil degradation, and viral defense. In the salivary proteome, 1,140 distinct proteins were detected. GSEA of GO Biological Processes in saliva during flare revealed significant enrichment for innate-immune pathways, including complement activation, neutrophil degranulation, and inflammatory response. A strong cross-modal agreement (r=0.6, Fig 1) was observed between the systemic transcriptomics and salivary proteomics, with 964 highly concordant pathways identified, notably high-level themes like infection response and immune response (Fig 2). Systemic transcriptomics had higher enrichment scores for these immune-related pathways (Fig 3).
Conclusion: The salivary proteome demonstrates marked immune activation and dysregulation of innate immunity in the oral mucosa during a PFAPA flare, closely mirroring findings in the systemic circulation. The striking inflammatory biological pathways enriched in the salivary proteome suggest that non-invasive salivary molecular profiling can effectively decipher PFAPA pathogenesis and offers a promising avenue for the future development of predictive biomarkers for PFAPA flares.
Figure 1: Correlation Matrix of Pathway Enrichment Metrics across RNA-seq and Proteomics Datasets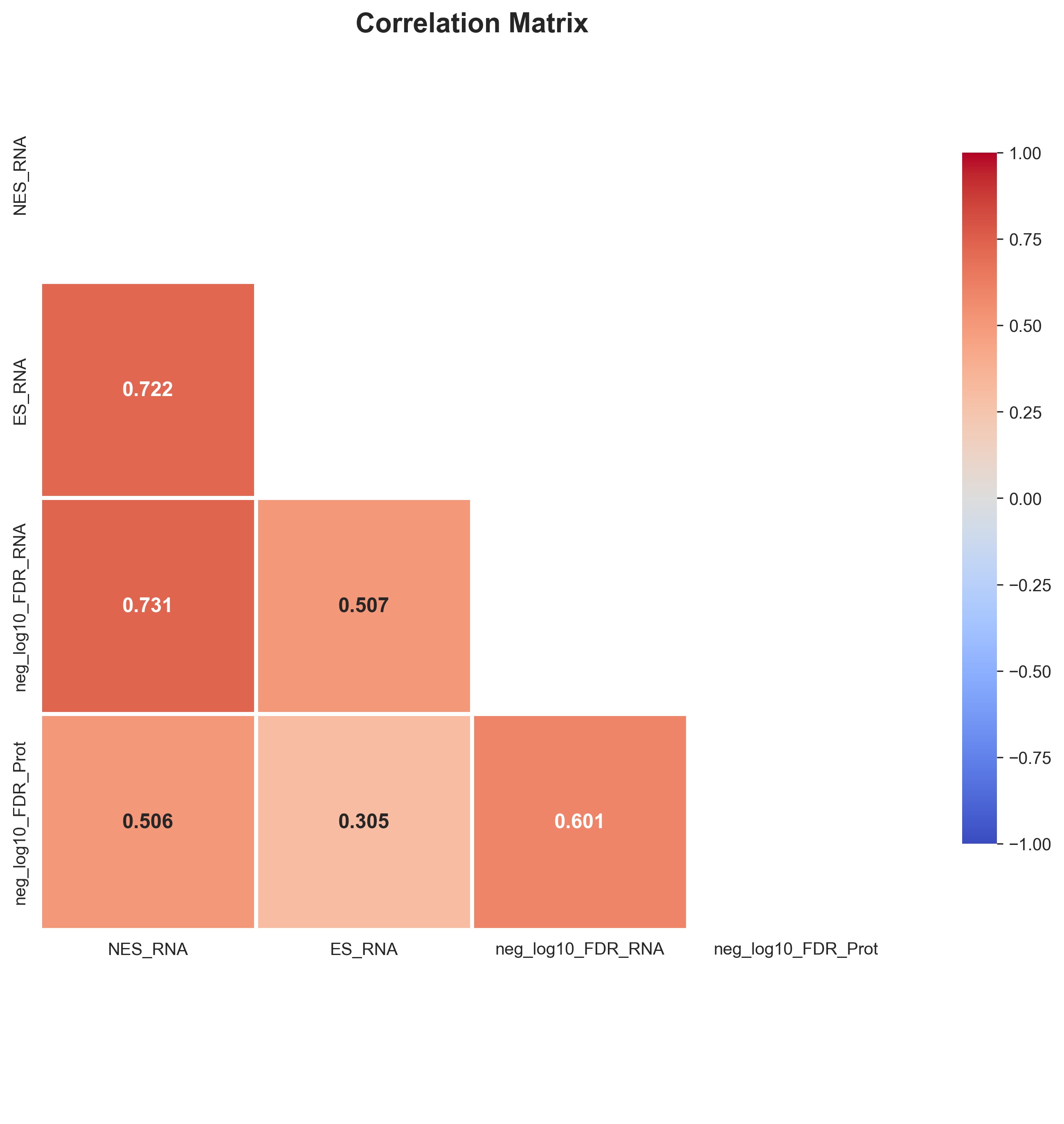 Correlation heatmap illustrating pairwise Pearson correlations among normalized enrichment scores (NES), enrichment scores (ES), and –log10(FDR) values from RNA-seq and proteomics analyses. Correlations are color-coded with positive correlations in red and negative in blue.
Correlation heatmap illustrating pairwise Pearson correlations among normalized enrichment scores (NES), enrichment scores (ES), and –log10(FDR) values from RNA-seq and proteomics analyses. Correlations are color-coded with positive correlations in red and negative in blue.
Figure 2: Top 20 Pathways by Agreement Score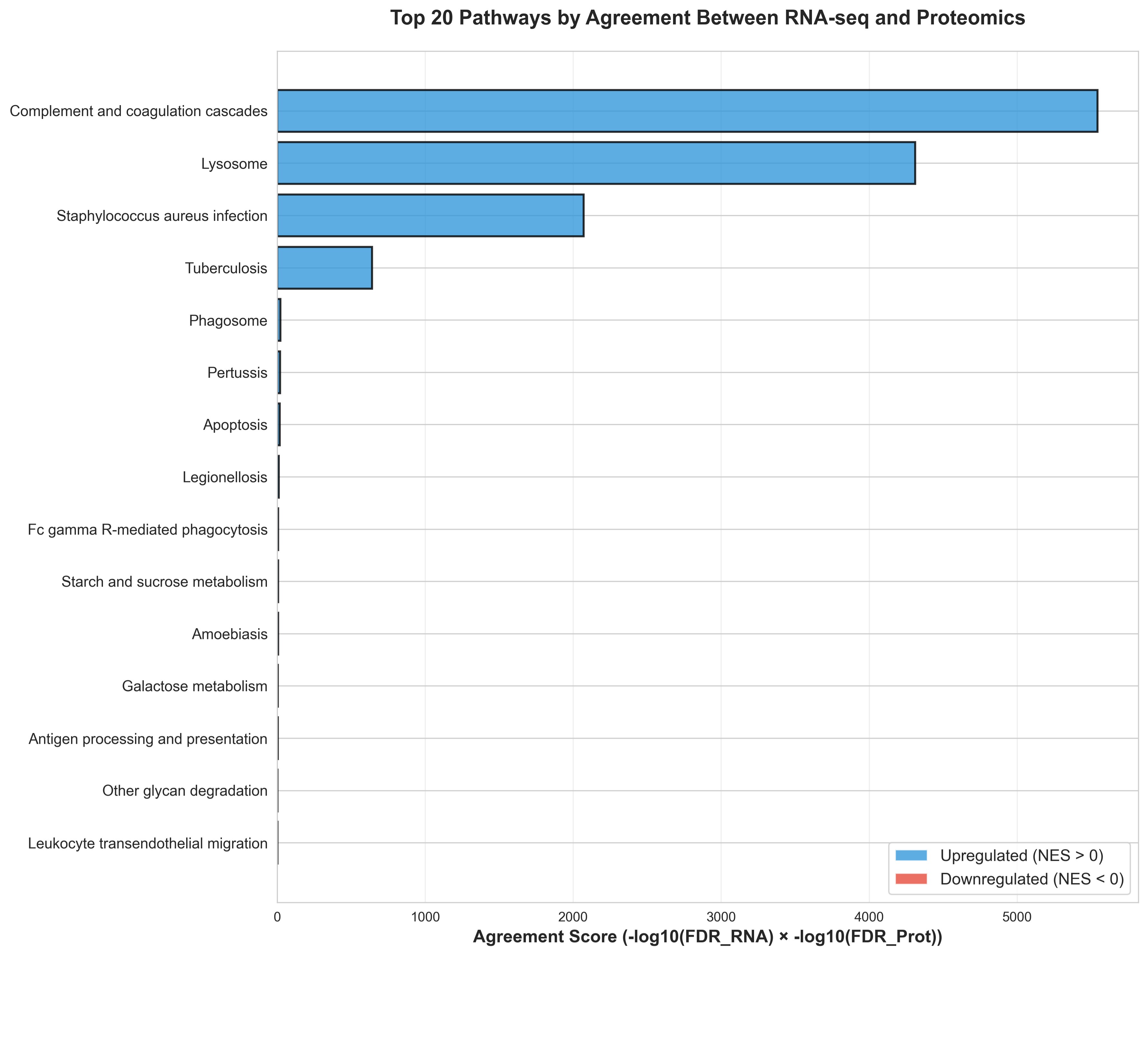 Bar plot showing the top 20 pathways ranked by an agreement score ( –log10(FDR) from RNA-seq x -log10(FDR) from proteomics). Higher scores indicate stronger concordance in pathway-level significance across the two omics layers. Bars are colored by the direction of regulation, with blue representing upregulated pathways (NES > 0) and red representing downregulated pathways (NES < 0).
Bar plot showing the top 20 pathways ranked by an agreement score ( –log10(FDR) from RNA-seq x -log10(FDR) from proteomics). Higher scores indicate stronger concordance in pathway-level significance across the two omics layers. Bars are colored by the direction of regulation, with blue representing upregulated pathways (NES > 0) and red representing downregulated pathways (NES < 0).
Figure 3 – Top 10 Enriched Pathways Identified by RNA-seq and Proteomics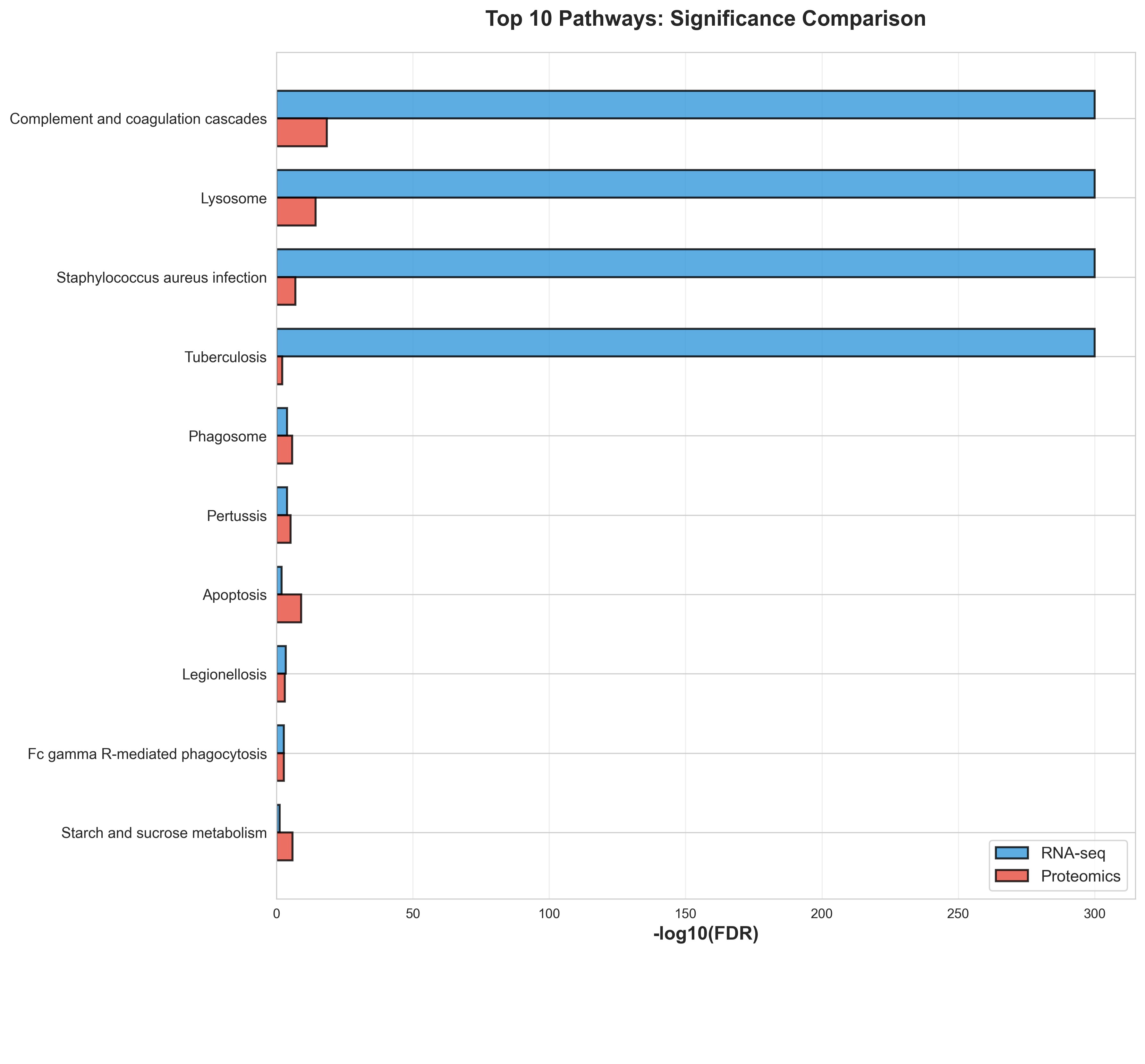 Bar plot showing the –log10(FDR) significance values for the top 10 enriched pathways from RNA-seq (blue) and proteomics (red) analyses. Pathways are ordered by significance. Bars of similar length indicate good agreement, while large differences suggest one dataset shows stronger enrichment than the other.
Bar plot showing the –log10(FDR) significance values for the top 10 enriched pathways from RNA-seq (blue) and proteomics (red) analyses. Pathways are ordered by significance. Bars of similar length indicate good agreement, while large differences suggest one dataset shows stronger enrichment than the other.
To cite this abstract in AMA style:
Lapidus S, Lozy T, Hakim E, Ambooken T, Weiss J, Li S, Golalipour E, Titova E, Adonimohammed S, Aalemansour H, Rodowicz A, Lejtman A, Haugh J, Sebbag A, Shivarathri R, Aptekmann A, Desai J. Unravelling the Pathogenesis of PFAPA (Periodic Fever, Aphthous Stomatitis, Pharyngitis and cervical Adenitis) Flares through Proteomics in the Salivary Matrix: A Potentially Non-Invasive Milieu for Biomarker Discovery [abstract]. Arthritis Rheumatol. 2026; 78 (suppl 3). https://acrabstracts.org/abstract/unravelling-the-pathogenesis-of-pfapa-periodic-fever-aphthous-stomatitis-pharyngitis-and-cervical-adenitis-flares-through-proteomics-in-the-salivary-matrix-a-potentially-non-invasive-milieu-for-b/. Accessed .« Back to 2026 Pediatric Rheumatology Symposium
ACR Meeting Abstracts - https://acrabstracts.org/abstract/unravelling-the-pathogenesis-of-pfapa-periodic-fever-aphthous-stomatitis-pharyngitis-and-cervical-adenitis-flares-through-proteomics-in-the-salivary-matrix-a-potentially-non-invasive-milieu-for-b/
