Session Information
Session Time: 6:00PM-7:00PM
Background/Purpose: As a multisystem vasculopathy and inflammatory myopathy, Juvenile dermatomyositis (JDM) is characterized by proximal muscle weakness, distinct rash, and risk of long-term complications such as calcinosis, skin ulceration, and chronic disease. Although Type I and Type II Interferon signatures are well established, this, coupled with myositis specific autoantibodies (MSAs), so far has not illustrated a complete understanding of the basic biology of this disease. Each unique MSA is associated with a unique clinical phenotype, although not every patient with a particular MSA demonstrates all the associated characteristics. Heterogeneity in outcomes necessitates a more fundamental understanding of disease mechanisms.
Our objective is to understand genome-scale gene dysregulation in JDM through a multiomic study of single nuclei from treatment naïve muscle biopsies, stratified by MSA. We hypothesize that distinct MSAs drive unique, cell-type-specific transcriptional and epigenomic programs in JDM.
Methods: Muscle biopsies were collected from treatment naïve patients diagnosed with JDM by the Bohan and Peter criteria in patients ≤18 years old, and congenital myopathy (CM). Biopsies were flash-frozen in liquid nitrogen and stored at -80°C (Cincinnati Children’s Hospital Medical Center (CCHMC) Pathology Core). Single nuclei isolation was performed (10x Genomics protocol with optimization performed on murine samples), followed by Chromium Next GEM Single Cell Multiome ATAC+ Gene Expression processing (CCHMC Single Cell Genomics Core).
Sequencing data were analyzed through DNA and RNA read alignment and quantification using CellRanger. Ambient mRNA was removed, doublets identified, and low-quality nuclei removed using CellBender. Data were visualized in a two-dimensional space using dimensionality reduction and cluster identification using Seurat.
Results: Single nuclei clustering revealed typical cell types consistent with previously demonstrated normal muscle tissue (Figure 1). Differentially expressed genes within individual cell types highlighted the known interferon signatures in multiple cell types (previously shown). Interestingly, we identified a subset of Type I Myonuclei which overexpress the CHD4 gene, only found in the biopsy from a patient with a Mi-2 MSA, and not found in such abundance in the other JDM samples, nor the CM samples (Figure 2).
Conclusion: We conducted a multiome study of single nuclei from treatment naïve muscle biopsies in JDM patients with a comparator group of congenital myopathy patients. While typical cell types of muscle are maintained, a unique cell population emerged from a patient with a Mi-2 MSA overexpressing the CHD4 gene, which is the biological target of the Mi-2 antibody. This observation, although preliminary and only tested and demonstrated in one sample so far, opens doors for further studies into the molecular dysfunction driving JDM pathology.
Figure 1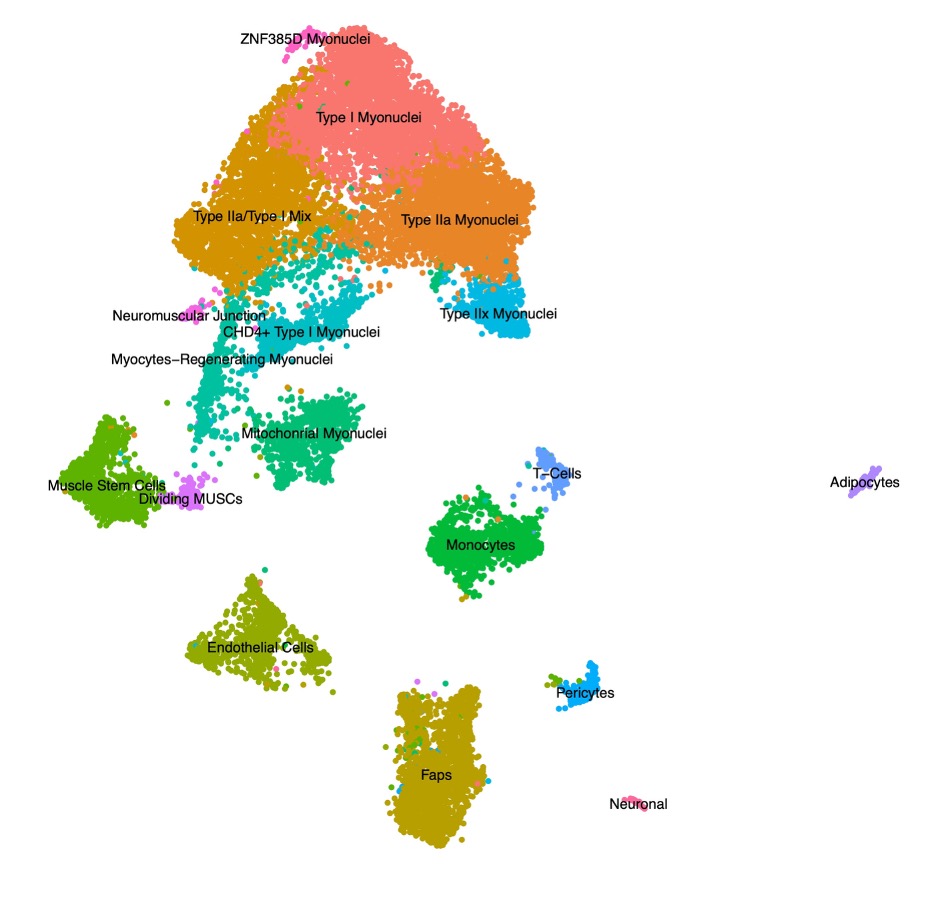 Figure 1. Uniform Manifold Approximation and Projection (UMAP) visualization represents sixteen color-coded nuclei clusters identified by tandem single nuclei RNA-sequencing and ATAC-sequencing. Data were generated from treatment naïve muscle biopsies from patients with juvenile dermatomyositis and congenital myopathy. FAPs = fibro-adipogenic precursors. MT = mitochondrial. NK = natural killer.
Figure 1. Uniform Manifold Approximation and Projection (UMAP) visualization represents sixteen color-coded nuclei clusters identified by tandem single nuclei RNA-sequencing and ATAC-sequencing. Data were generated from treatment naïve muscle biopsies from patients with juvenile dermatomyositis and congenital myopathy. FAPs = fibro-adipogenic precursors. MT = mitochondrial. NK = natural killer.
Figure 2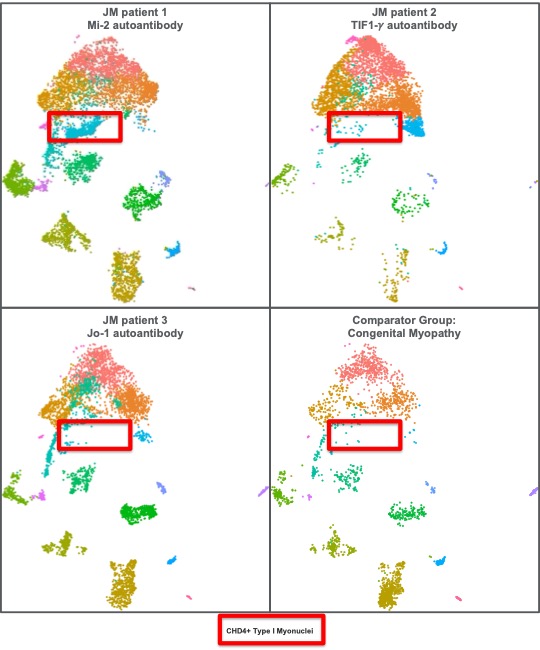 Figure 2. Uniform Manifold Approximation and Projection (UMAP) visualization from Figure 1, separated into individual patient samples with unique myositis specific antibodies, as well as a comparator group. FAPs = fibro-adipogenic precursors. MT = mitochondrial. NK = natural killer.
Figure 2. Uniform Manifold Approximation and Projection (UMAP) visualization from Figure 1, separated into individual patient samples with unique myositis specific antibodies, as well as a comparator group. FAPs = fibro-adipogenic precursors. MT = mitochondrial. NK = natural killer.
To cite this abstract in AMA style:
O'Connor S, Swoboda C, Weirauch M, Zygmunt A, Millay D, Kottyan L. Preliminary Findings: Single Nuclei RNAseq of Juvenile Dermatomyositis Muscle Biopsies Suggests Unique Myositis Specific Autoantibody Cell Population [abstract]. Arthritis Rheumatol. 2026; 78 (suppl 3). https://acrabstracts.org/abstract/preliminary-findings-single-nuclei-rnaseq-of-juvenile-dermatomyositis-muscle-biopsies-suggests-unique-myositis-specific-autoantibody-cell-population/. Accessed .« Back to 2026 Pediatric Rheumatology Symposium
ACR Meeting Abstracts - https://acrabstracts.org/abstract/preliminary-findings-single-nuclei-rnaseq-of-juvenile-dermatomyositis-muscle-biopsies-suggests-unique-myositis-specific-autoantibody-cell-population/
